[English] 日本語
 Yorodumi
Yorodumi- EMDB-13066: Metabolon-embedded pyruvate dehydrogenase complex E2 core at near... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13066 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Metabolon-embedded pyruvate dehydrogenase complex E2 core at near-atomic resolution | |||||||||
 Map data Map data | Icosahedral symmetrized map for the E2 core of PDHc from C. thermophilum | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | pyruvate / dehydrogenase / complex / e2 / core / c.thermophilum / metabolon / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationdihydrolipoyllysine-residue acetyltransferase / dihydrolipoyllysine-residue acetyltransferase activity / pyruvate decarboxylation to acetyl-CoA / pyruvate dehydrogenase complex / mitochondrion Similarity search - Function | |||||||||
| Biological species |  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) / Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) /  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) | |||||||||
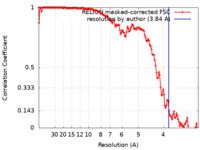
| Method | single particle reconstruction / cryo EM / Resolution: 3.84 Å | |||||||||
 Authors Authors | Tueting C / Kyrilis FL / Hamdi F / Kastritis PL | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Cryo-EM snapshots of a native lysate provide structural insights into a metabolon-embedded transacetylase reaction. Authors: Christian Tüting / Fotis L Kyrilis / Johannes Müller / Marija Sorokina / Ioannis Skalidis / Farzad Hamdi / Yashar Sadian / Panagiotis L Kastritis /   Abstract: Found across all kingdoms of life, 2-keto acid dehydrogenase complexes possess prominent metabolic roles and form major regulatory sites. Although their component structures are known, their higher- ...Found across all kingdoms of life, 2-keto acid dehydrogenase complexes possess prominent metabolic roles and form major regulatory sites. Although their component structures are known, their higher-order organization is highly heterogeneous, not only across species or tissues but also even within a single cell. Here, we report a cryo-EM structure of the fully active Chaetomium thermophilum pyruvate dehydrogenase complex (PDHc) core scaffold at 3.85 Å resolution (FSC = 0.143) from native cell extracts. By combining cryo-EM with macromolecular docking and molecular dynamics simulations, we resolve all PDHc core scaffold interfaces and dissect the residing transacetylase reaction. Electrostatics attract the lipoyl domain to the transacetylase active site and stabilize the coenzyme A, while apolar interactions position the lipoate in its binding cleft. Our results have direct implications on the structural determinants of the transacetylase reaction and the role of flexible regions in the context of the overall 10 MDa PDHc metabolon architecture. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13066.map.gz emd_13066.map.gz | 201.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13066-v30.xml emd-13066-v30.xml emd-13066.xml emd-13066.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13066_fsc.xml emd_13066_fsc.xml | 13.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_13066.png emd_13066.png | 200.6 KB | ||
| Masks |  emd_13066_msk_1.map emd_13066_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13066.cif.gz emd-13066.cif.gz | 6.2 KB | ||
| Others |  emd_13066_additional_1.map.gz emd_13066_additional_1.map.gz emd_13066_additional_2.map.gz emd_13066_additional_2.map.gz emd_13066_half_map_1.map.gz emd_13066_half_map_1.map.gz emd_13066_half_map_2.map.gz emd_13066_half_map_2.map.gz | 201 MB 200.7 MB 170.7 MB 170.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13066 http://ftp.pdbj.org/pub/emdb/structures/EMD-13066 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13066 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13066 | HTTPS FTP |
-Related structure data
| Related structure data |  7ottMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10892 (Title: Cryo-EM SPA dataset of Megadalton-range protein communities from a Chaetomium thermophilum native cell extract EMPIAR-10892 (Title: Cryo-EM SPA dataset of Megadalton-range protein communities from a Chaetomium thermophilum native cell extractData size: 1.1 TB Data #1: Unaligned fractions saved by Falcon 3 EC camera [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13066.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13066.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral symmetrized map for the E2 core of PDHc from C. thermophilum | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.5678 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_13066_msk_1.map emd_13066_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Asymmetric map for the E2 core of PDHc from C. thermophilum
| File | emd_13066_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Asymmetric map for the E2 core of PDHc from C. thermophilum | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: C2 symmetrized map for the E2 core of PDHc from C. thermophilum
| File | emd_13066_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C2 symmetrized map for the E2 core of PDHc from C. thermophilum | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Icosahedral symmetrized map for the E2 core of...
| File | emd_13066_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral symmetrized map for the E2 core of PDHc from C. thermophilum - Half-map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Icosahedral symmetrized map for the E2 core of...
| File | emd_13066_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Icosahedral symmetrized map for the E2 core of PDHc from C. thermophilum - Half-map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Metabolon-embedded pyruvate dehydrogenase complex E2 core at near...
| Entire | Name: Metabolon-embedded pyruvate dehydrogenase complex E2 core at near-atomic resolution |
|---|---|
| Components |
|
-Supramolecule #1: Metabolon-embedded pyruvate dehydrogenase complex E2 core at near...
| Supramolecule | Name: Metabolon-embedded pyruvate dehydrogenase complex E2 core at near-atomic resolution type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Icosahedrally symmetrized E2 core component of the pyruvate dehydrogenase complex metabolon from the thermophilic fungus Chaetomium thermophilum |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) |
-Macromolecule #1: Acetyltransferase component of pyruvate dehydrogenase complex
| Macromolecule | Name: Acetyltransferase component of pyruvate dehydrogenase complex type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: dihydrolipoyllysine-residue acetyltransferase |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 48.777488 KDa |
| Sequence | String: MLAQVLRRQA LQHVRLARAA APSLTRWYAS YPPHTIVKMP ALSPTMTSGN IGAWQKKPGD AITPGEVLVE IETDKAQMDF EFQEEGVLA KILKETGEKD VAVGSPIAVL VEEGTDINAF QNFTLEDAGG DAAAPAAPAK EELAKAETAP TPASTSAPEP E ETTSTGKL ...String: MLAQVLRRQA LQHVRLARAA APSLTRWYAS YPPHTIVKMP ALSPTMTSGN IGAWQKKPGD AITPGEVLVE IETDKAQMDF EFQEEGVLA KILKETGEKD VAVGSPIAVL VEEGTDINAF QNFTLEDAGG DAAAPAAPAK EELAKAETAP TPASTSAPEP E ETTSTGKL EPALDREPNV SFAAKKLAHE LDVPLKALKG TGPGGKITEE DVKKAASAPA AAAAAPGAAY QDIPISNMRK TI ATRLKES VSENPHFFVT SELSVSKLLK LRQALNSSAE GRYKLSVNDF LIKAIAVACK RVPAVNSSWR DGVIRQFDTV DVS VAVATP TGLITPIVKG VEAKGLETIS ATVKELAKKA RDGKLKPEDY QGGTISISNM GMNPAVERFT AIINPPQAAI LAVG TTKKV AVPVENEDGT TGVEWDDQIV VTASFDHKVV DGAVGAEWMR ELKKVVENPL ELLL UniProtKB: Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 200.0 mM / Component - Formula: C2H7NO2 / Component - Name: Ammonium ethanoate Details: Buffer was freshly made from solid ammonium acetate, filtrated, and degassed by ultrasonication. |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Pressure: 0.04 kPa |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Temperature | Min: 77.15 K / Max: 103.15 K |
| Alignment procedure | Coma free - Residual tilt: 14.7 mrad |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 2808 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 95677 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 92000 |
| Sample stage | Specimen holder model: OTHER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | The initial model was fitted using ChimeraX and then subsequently refined using iterative cycles of manual refinement using Coot and automatic Real-space refinement using PHENIX. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-7ott: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)