[English] 日本語
 Yorodumi
Yorodumi- EMDB-11951: Context-specific inhibition of eukaryotic translation by macrolid... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11951 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Context-specific inhibition of eukaryotic translation by macrolide antibiotics | |||||||||
 Map data Map data | Focused refinement for reconstruction of S. cervisiea ribosome with telithromycin at a final resolution of 2.9 A | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | context-specific inhibition / macrolide antibiotic / ribosome / telithromycin | |||||||||
| Function / homology |  Function and homology information Function and homology informationhexon binding / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / response to cycloheximide / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Formation of a pool of free 40S subunits / preribosome, large subunit precursor / L13a-mediated translational silencing of Ceruloplasmin expression ...hexon binding / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / response to cycloheximide / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Formation of a pool of free 40S subunits / preribosome, large subunit precursor / L13a-mediated translational silencing of Ceruloplasmin expression / translational elongation / ribosomal large subunit export from nucleus / regulation of translational fidelity / protein-RNA complex assembly / translational termination / maturation of LSU-rRNA / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal large subunit biogenesis / translational initiation / macroautophagy / maintenance of translational fidelity / modification-dependent protein catabolic process / rRNA processing / protein tag activity / ribosome biogenesis / viral capsid / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / 5S rRNA binding / cytoplasmic translation / cytosolic large ribosomal subunit / negative regulation of translation / rRNA binding / ribosome / protein ubiquitination / structural constituent of ribosome / translation / response to antibiotic / mRNA binding / ubiquitin protein ligase binding / host cell nucleus / nucleolus / RNA binding / nucleus / metal ion binding / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
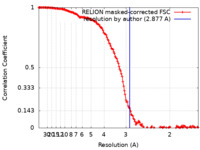
| Method | single particle reconstruction / cryo EM / Resolution: 2.877 Å | |||||||||
 Authors Authors | Koller TO / Wilson DN | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Context-specific action of macrolide antibiotics on the eukaryotic ribosome. Authors: Maxim S Svetlov / Timm O Koller / Sezen Meydan / Vaishnavi Shankar / Dorota Klepacki / Norbert Polacek / Nicholas R Guydosh / Nora Vázquez-Laslop / Daniel N Wilson / Alexander S Mankin /    Abstract: Macrolide antibiotics bind in the nascent peptide exit tunnel of the bacterial ribosome and prevent polymerization of specific amino acid sequences, selectively inhibiting translation of a subset of ...Macrolide antibiotics bind in the nascent peptide exit tunnel of the bacterial ribosome and prevent polymerization of specific amino acid sequences, selectively inhibiting translation of a subset of proteins. Because preventing translation of individual proteins could be beneficial for the treatment of human diseases, we asked whether macrolides, if bound to the eukaryotic ribosome, would retain their context- and protein-specific action. By introducing a single mutation in rRNA, we rendered yeast Saccharomyces cerevisiae cells sensitive to macrolides. Cryo-EM structural analysis showed that the macrolide telithromycin binds in the tunnel of the engineered eukaryotic ribosome. Genome-wide analysis of cellular translation and biochemical studies demonstrated that the drug inhibits eukaryotic translation by preferentially stalling ribosomes at distinct sequence motifs. Context-specific action markedly depends on the macrolide structure. Eliminating macrolide-arrest motifs from a protein renders its translation macrolide-tolerant. Our data illuminate the prospects of adapting macrolides for protein-selective translation inhibition in eukaryotic cells. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11951.map.gz emd_11951.map.gz | 265.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11951-v30.xml emd-11951-v30.xml emd-11951.xml emd-11951.xml | 61.9 KB 61.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11951_fsc.xml emd_11951_fsc.xml | 14.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_11951.png emd_11951.png | 44.4 KB | ||
| Filedesc metadata |  emd-11951.cif.gz emd-11951.cif.gz | 13.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11951 http://ftp.pdbj.org/pub/emdb/structures/EMD-11951 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11951 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11951 | HTTPS FTP |
-Validation report
| Summary document |  emd_11951_validation.pdf.gz emd_11951_validation.pdf.gz | 736.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11951_full_validation.pdf.gz emd_11951_full_validation.pdf.gz | 736.2 KB | Display | |
| Data in XML |  emd_11951_validation.xml.gz emd_11951_validation.xml.gz | 14.5 KB | Display | |
| Data in CIF |  emd_11951_validation.cif.gz emd_11951_validation.cif.gz | 19.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11951 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11951 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11951 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11951 | HTTPS FTP |
-Related structure data
| Related structure data |  7azyMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11951.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11951.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Focused refinement for reconstruction of S. cervisiea ribosome with telithromycin at a final resolution of 2.9 A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : 60S ribosomal subunit with 25S rRNA mutation G2400A and telithrom...
+Supramolecule #1: 60S ribosomal subunit with 25S rRNA mutation G2400A and telithrom...
+Macromolecule #1: 60S ribosomal protein L8-A
+Macromolecule #2: 60S ribosomal protein L23-A
+Macromolecule #3: 60S ribosomal protein L36-A
+Macromolecule #4: 60S ribosomal protein L9-A
+Macromolecule #5: 60S ribosomal protein L24-A
+Macromolecule #6: 60S ribosomal protein L37-A
+Macromolecule #7: 60S ribosomal protein L11-B
+Macromolecule #8: 60S ribosomal protein L25
+Macromolecule #9: 60S ribosomal protein L38
+Macromolecule #10: 60S ribosomal protein L13-A
+Macromolecule #11: 60S ribosomal protein L26-A
+Macromolecule #12: 60S ribosomal protein L39
+Macromolecule #13: 60S ribosomal protein L14-A
+Macromolecule #14: 60S ribosomal protein L27-A
+Macromolecule #15: Ubiquitin-60S ribosomal protein L40
+Macromolecule #16: 60S ribosomal protein L42-A
+Macromolecule #17: 60S ribosomal protein L15-A
+Macromolecule #18: 60S ribosomal protein L28
+Macromolecule #19: 60S ribosomal protein L43-A
+Macromolecule #20: 60S ribosomal protein L16-A
+Macromolecule #21: 60S ribosomal protein L29
+Macromolecule #22: 60S ribosomal protein L2-A
+Macromolecule #23: 60S ribosomal protein L17-A
+Macromolecule #24: 60S ribosomal protein L3
+Macromolecule #25: 60S ribosomal protein L18-A
+Macromolecule #26: 60S ribosomal protein L31-A
+Macromolecule #27: 60S ribosomal protein L10
+Macromolecule #28: 60S ribosomal protein L4-A
+Macromolecule #29: 60S ribosomal protein L32
+Macromolecule #30: 60S ribosomal protein L20-A
+Macromolecule #31: 60S ribosomal protein L5
+Macromolecule #32: 60S ribosomal protein L21-A
+Macromolecule #33: 60S ribosomal protein L33-A
+Macromolecule #34: 60S ribosomal protein L22-A
+Macromolecule #35: 60S ribosomal protein L6-A
+Macromolecule #36: 60S ribosomal protein L34-A
+Macromolecule #37: 60S ribosomal protein L7-A
+Macromolecule #38: 60S ribosomal protein L35-A
+Macromolecule #42: 60S ribosomal protein L19-A
+Macromolecule #39: 25S ribosomal RNA
+Macromolecule #40: 5S ribosomal RNA
+Macromolecule #41: 5.8S ribosomal RNA
+Macromolecule #43: TELITHROMYCIN
+Macromolecule #44: water
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R3/3 / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 3 / Pretreatment - Type: PLASMA CLEANING | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK III | ||||||||||||||||||
| Details | 5 Optical density at 260 nm. Approximatly 100 pmol of ribosome complex per grid. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 4371 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)