[English] 日本語
 Yorodumi
Yorodumi- EMDB-10284: Pseudomonas aeruginosa 70s ribosome from an aminoglycoside resist... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10284 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Pseudomonas aeruginosa 70s ribosome from an aminoglycoside resistant clinical isolate | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Ribosome / Pseudomonas aeruginosa | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationassembly of large subunit precursor of preribosome / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly ...assembly of large subunit precursor of preribosome / large ribosomal subunit / ribosomal small subunit assembly / transferase activity / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / RNA binding / metal ion binding / cytoplasm / cytosol Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.89 Å | |||||||||||||||
 Authors Authors | Halfon Y / Jimenez-Fernande A | |||||||||||||||
| Funding support |  Denmark, 4 items Denmark, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: Structure of ribosomes from an aminoglycoside-resistant clinical isolate. Authors: Yehuda Halfon / Alicia Jimenez-Fernandez / Ruggero La Rosa / Rocio Espinosa Portero / Helle Krogh Johansen / Donna Matzov / Zohar Eyal / Anat Bashan / Ella Zimmerman / Matthew Belousoff / ...Authors: Yehuda Halfon / Alicia Jimenez-Fernandez / Ruggero La Rosa / Rocio Espinosa Portero / Helle Krogh Johansen / Donna Matzov / Zohar Eyal / Anat Bashan / Ella Zimmerman / Matthew Belousoff / Søren Molin / Ada Yonath /    Abstract: Resistance to antibiotics has become a major threat to modern medicine. The ribosome plays a fundamental role in cell vitality by the translation of the genetic code into proteins; hence, it is a ...Resistance to antibiotics has become a major threat to modern medicine. The ribosome plays a fundamental role in cell vitality by the translation of the genetic code into proteins; hence, it is a major target for clinically useful antibiotics. We report here the cryo-electron microscopy structures of the ribosome of a pathogenic aminoglycoside (AG)-resistant strain, as well as of a nonresistance strain isolated from a cystic fibrosis patient. The structural studies disclosed defective ribosome complex formation due to a conformational change of rRNA helix H69, an essential intersubunit bridge, and a secondary binding site of the AGs. In addition, a stable conformation of nucleotides A1486 and A1487, pointing into helix h44, is created compared to a non-AG-bound ribosome. We suggest that altering the conformations of ribosomal protein uL6 and rRNA helix H69, which interact with initiation-factor IF2, interferes with proper protein synthesis initiation. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10284.map.gz emd_10284.map.gz | 28.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10284-v30.xml emd-10284-v30.xml emd-10284.xml emd-10284.xml | 66.7 KB 66.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10284.png emd_10284.png | 93.1 KB | ||
| Filedesc metadata |  emd-10284.cif.gz emd-10284.cif.gz | 12.9 KB | ||
| Others |  emd_10284_additional.map.gz emd_10284_additional.map.gz | 192.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10284 http://ftp.pdbj.org/pub/emdb/structures/EMD-10284 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10284 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10284 | HTTPS FTP |
-Related structure data
| Related structure data |  6spfMC  6spbC  6spcC  6spdC  6speC  6spgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10284.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10284.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
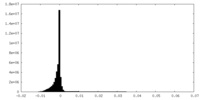
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Unsharp map
| File | emd_10284_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharp map | ||||||||||||
| Projections & Slices |
| ||||||||||||
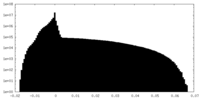
| Density Histograms |
- Sample components
Sample components
+Entire : Pseudomonas aeruginosa 70s ribosome from a clinical isolate
+Supramolecule #1: Pseudomonas aeruginosa 70s ribosome from a clinical isolate
+Macromolecule #1: Pseudomonas aeruginosa strain PAO1 23S ribosomal RNA
+Macromolecule #2: 5S rRNA
+Macromolecule #33: 16S rRNA
+Macromolecule #3: 50S ribosomal protein L2
+Macromolecule #4: 50S ribosomal protein L3
+Macromolecule #5: 50S ribosomal protein L4
+Macromolecule #6: 50S ribosomal protein L5
+Macromolecule #7: 50S ribosomal protein L6
+Macromolecule #8: 50S ribosomal protein L9
+Macromolecule #9: 50S ribosomal protein L11
+Macromolecule #10: 50S ribosomal protein L13
+Macromolecule #11: 50S ribosomal protein L14
+Macromolecule #12: 50S ribosomal protein L15
+Macromolecule #13: 50S ribosomal protein L16
+Macromolecule #14: 50S ribosomal protein L17
+Macromolecule #15: 50S ribosomal protein L18
+Macromolecule #16: 50S ribosomal protein L19
+Macromolecule #17: 50S ribosomal protein L20
+Macromolecule #18: 50S ribosomal protein L21
+Macromolecule #19: 50S ribosomal protein L22
+Macromolecule #20: 50S ribosomal protein L23
+Macromolecule #21: 50S ribosomal protein L24
+Macromolecule #22: 50S ribosomal protein L25
+Macromolecule #23: 50S ribosomal protein L27
+Macromolecule #24: 50S ribosomal protein L28
+Macromolecule #25: Ribosomal protein uL29
+Macromolecule #26: 50S ribosomal protein L30
+Macromolecule #27: 50S ribosomal protein L31
+Macromolecule #28: 50S ribosomal protein L32
+Macromolecule #29: 50S ribosomal protein L33
+Macromolecule #30: 50S ribosomal protein L34
+Macromolecule #31: 50S ribosomal protein L35
+Macromolecule #32: 50S ribosomal protein L36
+Macromolecule #34: 30S ribosomal protein S2
+Macromolecule #35: 30S ribosomal protein S3
+Macromolecule #36: 30S ribosomal protein S4
+Macromolecule #37: 30S ribosomal protein S5
+Macromolecule #38: 30S ribosomal protein S6
+Macromolecule #39: 30S ribosomal protein S7
+Macromolecule #40: 30S ribosomal protein S8
+Macromolecule #41: 30S ribosomal protein S9
+Macromolecule #42: 30S ribosomal protein S10
+Macromolecule #43: 30S ribosomal protein S11
+Macromolecule #44: 30S ribosomal protein S12
+Macromolecule #45: 30S ribosomal protein S13
+Macromolecule #46: 30S ribosomal protein S14
+Macromolecule #47: 30S ribosomal protein S15
+Macromolecule #48: 30S ribosomal protein S16
+Macromolecule #49: 30S ribosomal protein S17
+Macromolecule #50: 30S ribosomal protein S18
+Macromolecule #51: 30S ribosomal protein S19
+Macromolecule #52: 30S ribosomal protein S20
+Macromolecule #53: 30S ribosomal protein S21
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)