[English] 日本語
 Yorodumi
Yorodumi- EMDB-11827: Nonameric cytoplasmic domain of FlhA from Vibrio parahaemolyticus -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11827 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
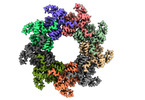
| Title | Nonameric cytoplasmic domain of FlhA from Vibrio parahaemolyticus | ||||||||||||||||||
 Map data Map data | postprocessed map | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | T3SS / export apparatus / secretion / protein export / motility / PROTEIN TRANSPORT | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum assembly / protein secretion / membrane => GO:0016020 / plasma membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
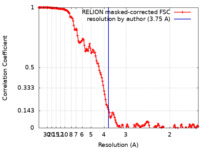
| Method | single particle reconstruction / cryo EM / Resolution: 3.75 Å | ||||||||||||||||||
 Authors Authors | Kuhlen L / Johnson S | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: PLoS One / Year: 2021 Journal: PLoS One / Year: 2021Title: Nonameric structures of the cytoplasmic domain of FlhA and SctV in the context of the full-length protein. Authors: Lucas Kuhlen / Steven Johnson / Jerry Cao / Justin C Deme / Susan M Lea /   Abstract: Type three secretion is the mechanism of protein secretion found in bacterial flagella and injectisomes. At its centre is the export apparatus (EA), a complex of five membrane proteins through which ...Type three secretion is the mechanism of protein secretion found in bacterial flagella and injectisomes. At its centre is the export apparatus (EA), a complex of five membrane proteins through which secretion substrates pass the inner membrane. While the complex formed by four of the EA proteins has been well characterised structurally, little is known about the structure of the membrane domain of the largest subunit, FlhA in flagella, SctV in injectisomes. Furthermore, the biologically relevant nonameric assembly of FlhA/SctV has been infrequently observed and differences in conformation of the cytoplasmic portion of FlhA/SctV between open and closed states have been suggested to reflect secretion system specific differences. FlhA has been shown to bind to chaperone-substrate complexes in an open state, but in previous assembled ring structures, SctV is in a closed state. Here, we identify FlhA and SctV homologues that can be recombinantly produced in the oligomeric state and study them using cryo-electron microscopy. The structures of the cytoplasmic domains from both FlhA and SctV are in the open state and we observe a conserved interaction between a short stretch of residues at the N-terminus of the cytoplasmic domain, known as FlhAL/SctVL, with a groove on the adjacent protomer's cytoplasmic domain, which stabilises the nonameric ring assembly. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11827.map.gz emd_11827.map.gz | 265.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11827-v30.xml emd-11827-v30.xml emd-11827.xml emd-11827.xml | 15.4 KB 15.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11827_fsc.xml emd_11827_fsc.xml | 15.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_11827.png emd_11827.png | 191.2 KB | ||
| Masks |  emd_11827_msk_1.map emd_11827_msk_1.map | 290.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-11827.cif.gz emd-11827.cif.gz | 5.7 KB | ||
| Others |  emd_11827_half_map_1.map.gz emd_11827_half_map_1.map.gz emd_11827_half_map_2.map.gz emd_11827_half_map_2.map.gz | 231.8 MB 231.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11827 http://ftp.pdbj.org/pub/emdb/structures/EMD-11827 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11827 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11827 | HTTPS FTP |
-Validation report
| Summary document |  emd_11827_validation.pdf.gz emd_11827_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11827_full_validation.pdf.gz emd_11827_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_11827_validation.xml.gz emd_11827_validation.xml.gz | 22.5 KB | Display | |
| Data in CIF |  emd_11827_validation.cif.gz emd_11827_validation.cif.gz | 29.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11827 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11827 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11827 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11827 | HTTPS FTP |
-Related structure data
| Related structure data |  7amyMC  7alwC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11827.map.gz / Format: CCP4 / Size: 290.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11827.map.gz / Format: CCP4 / Size: 290.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | postprocessed map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11827_msk_1.map emd_11827_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_11827_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_11827_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Nonamer of the FlhA cytoplasmic domain from Vibrio parahaemolyticus
| Entire | Name: Nonamer of the FlhA cytoplasmic domain from Vibrio parahaemolyticus |
|---|---|
| Components |
|
-Supramolecule #1: Nonamer of the FlhA cytoplasmic domain from Vibrio parahaemolyticus
| Supramolecule | Name: Nonamer of the FlhA cytoplasmic domain from Vibrio parahaemolyticus type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 700 KDa |
-Macromolecule #1: Flagellar biosynthesis protein FlhA
| Macromolecule | Name: Flagellar biosynthesis protein FlhA / type: protein_or_peptide / ID: 1 / Number of copies: 9 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 76.55582 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLTRLKQLQT STKGYIGIPI VLLMILAMVI LPLPPLLLDA LFTFNIVLAI LVLLVSTTAK RPLDFSVFPT ILLVATLLRL TLNVASTRI VLLEGHNGGD AAGKVIQAFG EVVIGGNYVV GMVVFIILMI INFVVITKGG ERISEVSARF TLDALPGKQM A IDADLNAG ...String: MLTRLKQLQT STKGYIGIPI VLLMILAMVI LPLPPLLLDA LFTFNIVLAI LVLLVSTTAK RPLDFSVFPT ILLVATLLRL TLNVASTRI VLLEGHNGGD AAGKVIQAFG EVVIGGNYVV GMVVFIILMI INFVVITKGG ERISEVSARF TLDALPGKQM A IDADLNAG LIDQETARLR RKEVANEADF HGSMDGASKF VRGDAVAGLL ILFINIIGGI SIGVFEHGLP ASEAFKTYAL LT IGDGLVA QIPSLLLATA AAIIVTRIND SDNGMSETMQ KQLLATPATL FTVAGIMAVI GMVPGMPHLA FFAFAGALGF AGW RQSKKP VQDTQIEQVE ALSQAMQEED TPLTWDDIPH VHTLSLALGY RLVHLVNKDQ GAPLSQRIRG VRRNLSEQVG FLLP EVRIR DNLSLKPNQY TISLNGEVIE QGFIEPERLM AIAVGDTYGE IDGILGSDPA YQLPAVWIEH QDKAKALNMG YQVVD DGTV IATHISKIMK TNLAELFTHD DVEAMTQRLT QQAPKLAEAL AAALNPAQQL KVYRQLLLDQ VPLKDIRTIA NTMLES SEN TKDPILLAAD VRCALKRTLV NLIAGQKPEL NVYALSDELE QMLLTSLQQA QASGTVVLDS FPIEPNILGQ FQQNLPL IR QQLKQQGLPP ILLVMPQLRP LLARYARTFT QGLAVLSYNE IPENKQINVV GNLGENLYFQ UniProtKB: Flagellar biosynthesis protein FlhA |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X