[English] 日本語
 Yorodumi
Yorodumi- EMDB-1138: Structure of the hepatitis C virus IRES bound to the human 80S ri... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1138 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the hepatitis C virus IRES bound to the human 80S ribosome: remodeling of the HCV IRES. | |||||||||
 Map data Map data | Cryo-EM map of the hepatitis C virus internal ribosome entry site in complex with human 80S ribosome | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Hepatitis C virus Hepatitis C virus | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 15.0 Å | |||||||||
 Authors Authors | Boehringer D / Thermann R / Ostareck-Lederer A / Lewis JD / Stark H | |||||||||
 Citation Citation |  Journal: Structure / Year: 2005 Journal: Structure / Year: 2005Title: Structure of the hepatitis C virus IRES bound to the human 80S ribosome: remodeling of the HCV IRES. Authors: Daniel Boehringer / Rolf Thermann / Antje Ostareck-Lederer / Joe D Lewis / Holger Stark /  Abstract: Initiation of translation of the hepatitis C virus (HCV) polyprotein is driven by an internal ribosome entry site (IRES) RNA that bypasses much of the eukaryotic translation initiation machinery. ...Initiation of translation of the hepatitis C virus (HCV) polyprotein is driven by an internal ribosome entry site (IRES) RNA that bypasses much of the eukaryotic translation initiation machinery. Here, single-particle electron cryomicroscopy has been used to study the mechanism of HCV IRES-mediated initiation. A HeLa in vitro translation system was used to assemble human IRES-80S ribosome complexes under near physiological conditions; these were stalled before elongation. Domain 2 of the HCV IRES is bound to the tRNA exit site, touching the L1 stalk of the 60S subunit, suggesting a mechanism for the removal of the HCV IRES in the progression to elongation. Domain 3 of the HCV IRES positions the initiation codon in the ribosomal mRNA binding cleft by binding helix 28 at the head of the 40S subunit. The comparison with the previously published binary 40S-HCV IRES complex reveals structural rearrangements in the two pseudoknot structures of the HCV IRES in translation initiation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1138.map.gz emd_1138.map.gz | 7.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1138-v30.xml emd-1138-v30.xml emd-1138.xml emd-1138.xml | 7.9 KB 7.9 KB | Display Display |  EMDB header EMDB header |
| Images |  1138.gif 1138.gif | 9.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1138 http://ftp.pdbj.org/pub/emdb/structures/EMD-1138 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1138 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1138 | HTTPS FTP |
-Related structure data
| Related structure data |  2agnMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_1138.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1138.map.gz / Format: CCP4 / Size: 7.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of the hepatitis C virus internal ribosome entry site in complex with human 80S ribosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.6 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
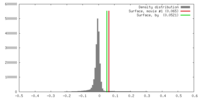
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : IRES-80S ribosome complex
| Entire | Name: IRES-80S ribosome complex |
|---|---|
| Components |
|
-Supramolecule #1000: IRES-80S ribosome complex
| Supramolecule | Name: IRES-80S ribosome complex / type: sample / ID: 1000 / Number unique components: 2 |
|---|
-Supramolecule #1: ribosome
| Supramolecule | Name: ribosome / type: organelle_or_cellular_component / ID: 1 / Details: H. sapiens / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: human / Cell: HeLa Homo sapiens (human) / synonym: human / Cell: HeLa |
-Macromolecule #1: HCV IRES
| Macromolecule | Name: HCV IRES / type: rna / ID: 1 / Classification: OTHER / Structure: DOUBLE HELIX / Synthetic?: Yes |
|---|---|
| Source (natural) | Organism:  Hepatitis C virus Hepatitis C virus |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.1 Details: 20 mM Tris.HCl pH 8.1, 145 mM KCl, 1 mM CaCl2, 5 mM MgCl2, 0.2 mM DTT, 5 mM tobramycin |
|---|---|
| Staining | Type: NEGATIVE / Details: native cryo |
| Grid | Details: copper grid |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Image recording | Category: CCD / Film or detector model: KODAK SO-163 FILM |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: OTHER |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 15.0 Å / Resolution method: FSC 3 SIGMA CUT-OFF / Software - Name: imagic 5 / Number images used: 24100 |
|---|
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)