[English] 日本語
 Yorodumi
Yorodumi- EMDB-10525: Cryo-EM structure of Toxoplasma gondii mitochondrial ATP synthase... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10525 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Toxoplasma gondii mitochondrial ATP synthase hexamer, membrane region | |||||||||||||||
 Map data Map data | Toxoplasma gondii ATP synthase hexamer, membrane region | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | mitochondrial / ATP synthase / hexamer / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationmitochondrial electron transport, ubiquinol to cytochrome c / proton transmembrane transporter activity / proton motive force-driven ATP synthesis / proton-transporting two-sector ATPase complex, proton-transporting domain / proton motive force-driven mitochondrial ATP synthesis / H+-transporting two-sector ATPase / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / ADP binding / electron transfer activity ...mitochondrial electron transport, ubiquinol to cytochrome c / proton transmembrane transporter activity / proton motive force-driven ATP synthesis / proton-transporting two-sector ATPase complex, proton-transporting domain / proton motive force-driven mitochondrial ATP synthesis / H+-transporting two-sector ATPase / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / ADP binding / electron transfer activity / mitochondrial inner membrane / hydrolase activity / heme binding / lipid binding / mitochondrion / ATP binding / membrane / metal ion binding Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
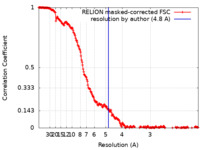
| Method | single particle reconstruction / cryo EM / Resolution: 4.8 Å | |||||||||||||||
 Authors Authors | Muhleip A / Kock Flygaard R / Amunts A | |||||||||||||||
| Funding support |  Sweden, 4 items Sweden, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: ATP synthase hexamer assemblies shape cristae of Toxoplasma mitochondria. Authors: Alexander Mühleip / Rasmus Kock Flygaard / Jana Ovciarikova / Alice Lacombe / Paula Fernandes / Lilach Sheiner / Alexey Amunts /   Abstract: Mitochondrial ATP synthase plays a key role in inducing membrane curvature to establish cristae. In Apicomplexa causing diseases such as malaria and toxoplasmosis, an unusual cristae morphology has ...Mitochondrial ATP synthase plays a key role in inducing membrane curvature to establish cristae. In Apicomplexa causing diseases such as malaria and toxoplasmosis, an unusual cristae morphology has been observed, but its structural basis is unknown. Here, we report that the apicomplexan ATP synthase assembles into cyclic hexamers, essential to shape their distinct cristae. Cryo-EM was used to determine the structure of the hexamer, which is held together by interactions between parasite-specific subunits in the lumenal region. Overall, we identified 17 apicomplexan-specific subunits, and a minimal and nuclear-encoded subunit-a. The hexamer consists of three dimers with an extensive dimer interface that includes bound cardiolipins and the inhibitor IF. Cryo-ET and subtomogram averaging revealed that hexamers arrange into ~20-megadalton pentagonal pyramids in the curved apical membrane regions. Knockout of the linker protein ATPTG11 resulted in the loss of pentagonal pyramids with concomitant aberrantly shaped cristae. Together, this demonstrates that the unique macromolecular arrangement is critical for the maintenance of cristae morphology in Apicomplexa. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10525.map.gz emd_10525.map.gz | 431.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10525-v30.xml emd-10525-v30.xml emd-10525.xml emd-10525.xml | 51.7 KB 51.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10525_fsc.xml emd_10525_fsc.xml | 21.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_10525.png emd_10525.png | 115.4 KB | ||
| Masks |  emd_10525_msk_1.map emd_10525_msk_1.map | 824 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10525.cif.gz emd-10525.cif.gz | 11.9 KB | ||
| Others |  emd_10525_half_map_1.map.gz emd_10525_half_map_1.map.gz emd_10525_half_map_2.map.gz emd_10525_half_map_2.map.gz | 664.4 MB 665.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10525 http://ftp.pdbj.org/pub/emdb/structures/EMD-10525 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10525 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10525 | HTTPS FTP |
-Related structure data
| Related structure data |  6tmlMC  6tmgC  6tmhC  6tmiC  6tmjC  6tmkC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10525.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10525.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Toxoplasma gondii ATP synthase hexamer, membrane region | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
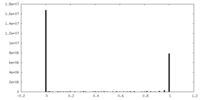
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10525_msk_1.map emd_10525_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 2
| File | emd_10525_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 1
| File | emd_10525_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Toxoplasma gondii ATP synthase hexamer
+Supramolecule #1: Toxoplasma gondii ATP synthase hexamer
+Macromolecule #1: ATPTG11
+Macromolecule #2: ATPTG7
+Macromolecule #3: ATPTG14
+Macromolecule #4: ATPTG5
+Macromolecule #5: subunit k
+Macromolecule #6: subunit a
+Macromolecule #7: subunit i/j
+Macromolecule #8: ATPTG13
+Macromolecule #9: ATPTG15
+Macromolecule #10: ATPTG6
+Macromolecule #11: ATPTG3
+Macromolecule #12: ATPTG17,ATPTG17,ATPTG17
+Macromolecule #13: subunit b
+Macromolecule #14: ATPTG12
+Macromolecule #15: ATPTG10
+Macromolecule #16: subunit f
+Macromolecule #17: ATPTG8
+Macromolecule #18: ATPTG1
+Macromolecule #19: ATPTG2
+Macromolecule #20: subunit 8
+Macromolecule #21: ATPTG9
+Macromolecule #22: ATPTG4
+Macromolecule #23: ATPTG16
+Macromolecule #24: subunit d
+Macromolecule #25: Oligomycin sensitivity conferring protein (OSCP)
+Macromolecule #26: Inhibitor of F1
+Macromolecule #27: ATP synthase subunit alpha
+Macromolecule #28: ATP synthase subunit beta
+Macromolecule #29: ATP synthase subunit gamma
+Macromolecule #30: ATP synthase subunit delta
+Macromolecule #31: ATP synthase subunit epsilon
+Macromolecule #32: subunit c
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 19 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Number real images: 7604 / Average electron dose: 32.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-6tml: |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)