+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0868 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
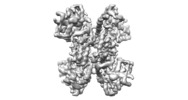
| Title | Cryo-EM structure of the AtMLKL2 tetramer | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Plant MLKLs / Cryo-EM / Tetramer / CYTOSOLIC PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationprotein serine/threonine/tyrosine kinase activity / protein kinase activity / signal transduction / ATP binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Lisa M / Huang M | |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Cryo-EM structure of the AtMLKL3 tetramer Authors: Lisa M / Huang M / Zhang X / Ryohei TN / Leila BK / Isabel MLS / Florence J / Viera K / Dmitry L / Jane EP / James MM / Kay H / Paul SL / Chai J / Takaki M | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0868.map.gz emd_0868.map.gz | 3.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0868-v30.xml emd-0868-v30.xml emd-0868.xml emd-0868.xml | 9.1 KB 9.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0868.png emd_0868.png | 79 KB | ||
| Filedesc metadata |  emd-0868.cif.gz emd-0868.cif.gz | 5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0868 http://ftp.pdbj.org/pub/emdb/structures/EMD-0868 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0868 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0868 | HTTPS FTP |
-Validation report
| Summary document |  emd_0868_validation.pdf.gz emd_0868_validation.pdf.gz | 389.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0868_full_validation.pdf.gz emd_0868_full_validation.pdf.gz | 389.4 KB | Display | |
| Data in XML |  emd_0868_validation.xml.gz emd_0868_validation.xml.gz | 5.7 KB | Display | |
| Data in CIF |  emd_0868_validation.cif.gz emd_0868_validation.cif.gz | 6.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0868 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0868 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0868 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0868 | HTTPS FTP |
-Related structure data
| Related structure data |  6lbaMC  9954C  6ka4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0868.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0868.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.30654 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mixed lineage kinase domain-like (MLKL) protein 2
| Entire | Name: Mixed lineage kinase domain-like (MLKL) protein 2 |
|---|---|
| Components |
|
-Supramolecule #1: Mixed lineage kinase domain-like (MLKL) protein 2
| Supramolecule | Name: Mixed lineage kinase domain-like (MLKL) protein 2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Protein kinase family protein
| Macromolecule | Name: Protein kinase family protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 66.682531 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: AAMEQFRQIG EVLGSLNALM VLQDDILINQ RQCCLLLELF SLAFNTVAEE IRQNLKLEEK HTKWRALEQP LRELTRVFKE GELYVKHCM DNSDWWGKVI NLHQNKDCVE FHIHNLFCYF SAVVEAIEAA GEISGLDPSE MERRRVVFSR KYDREWNDPK M FQWRFGKQ ...String: AAMEQFRQIG EVLGSLNALM VLQDDILINQ RQCCLLLELF SLAFNTVAEE IRQNLKLEEK HTKWRALEQP LRELTRVFKE GELYVKHCM DNSDWWGKVI NLHQNKDCVE FHIHNLFCYF SAVVEAIEAA GEISGLDPSE MERRRVVFSR KYDREWNDPK M FQWRFGKQ YLLSRDICSR FEHSWREDRW NLVEALQEKR KSDSDDIGKT EKRLADLLLK KLTGLEQFNG KLFPSSILLG SK DYQVKKR LDADGQYKEI QWLGDSFALR HFFSDLEPLS SEISSLLALC HSNILQYLCG FYDEERKECF LVMELMHKDL QSY MKENCG PRRRYLFSIP VVIDIMLQIA RGMEYLHGND IFHGDLNPMN IHLKERSHTE GYFHAKICGF GLSSVVKAQS SSKP GTPDP VIWYAPEVLA EMEQDLNGKT PKSKLTHKAD VYSFAMVCFE LITGKVPFED SHLQGEPMTI NIRMGERPLF PFPSP KTLV SLIKRCWHSE PSQRPNFSSI CRILRYIKKF LVVNPDHGHP QMQTPLVDCW DLEARFLRKF PGDAGSHTAS VNQIPF QLT SYRVLEKEK UniProtKB: Protein kinase family protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.5625 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.1 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 1527009 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller