+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0492 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | hTRiC-hPFD Class1 (No PFD) | ||||||||||||||||||||||||
 Map data Map data | hTRiC-hPFD Class1 (No PFD) | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | TRiC/CCT / PFD / CryoEM / Molecular Chaperone / Protein folding / CHAPERONE | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationscaRNA localization to Cajal body / zona pellucida receptor complex / positive regulation of establishment of protein localization to telomere / positive regulation of protein localization to Cajal body / chaperone mediated protein folding independent of cofactor / tubulin complex assembly / BBSome-mediated cargo-targeting to cilium / binding of sperm to zona pellucida / positive regulation of telomerase RNA localization to Cajal body / Folding of actin by CCT/TriC ...scaRNA localization to Cajal body / zona pellucida receptor complex / positive regulation of establishment of protein localization to telomere / positive regulation of protein localization to Cajal body / chaperone mediated protein folding independent of cofactor / tubulin complex assembly / BBSome-mediated cargo-targeting to cilium / binding of sperm to zona pellucida / positive regulation of telomerase RNA localization to Cajal body / Folding of actin by CCT/TriC / Formation of tubulin folding intermediates by CCT/TriC / chaperonin-containing T-complex / Prefoldin mediated transfer of substrate to CCT/TriC / RHOBTB1 GTPase cycle / intermediate filament cytoskeleton / WD40-repeat domain binding / pericentriolar material / beta-tubulin binding / Association of TriC/CCT with target proteins during biosynthesis / : / chaperone-mediated protein complex assembly / RHOBTB2 GTPase cycle / heterochromatin / chaperone-mediated protein folding / protein folding chaperone / positive regulation of telomere maintenance via telomerase / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / acrosomal vesicle / cell projection / mRNA 3'-UTR binding / ATP-dependent protein folding chaperone / response to virus / cilium / mRNA 5'-UTR binding / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / azurophil granule lumen / G-protein beta-subunit binding / unfolded protein binding / melanosome / protein folding / cell body / secretory granule lumen / ficolin-1-rich granule lumen / microtubule / cytoskeleton / protein stabilization / cadherin binding / centrosome / ubiquitin protein ligase binding / Neutrophil degranulation / Golgi apparatus / ATP hydrolysis activity / RNA binding / extracellular exosome / extracellular region / nucleoplasm / ATP binding / cytosol / cytoplasm Similarity search - Function | ||||||||||||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||||||||||||||
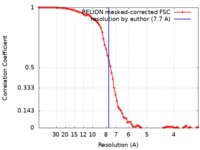
| Method | single particle reconstruction / cryo EM / Resolution: 7.7 Å | ||||||||||||||||||||||||
 Authors Authors | Gestaut DR / Roh SH | ||||||||||||||||||||||||
| Funding support |  United States, European Union, 7 items United States, European Union, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Cell / Year: 2019 Journal: Cell / Year: 2019Title: The Chaperonin TRiC/CCT Associates with Prefoldin through a Conserved Electrostatic Interface Essential for Cellular Proteostasis. Authors: Daniel Gestaut / Soung Hun Roh / Boxue Ma / Grigore Pintilie / Lukasz A Joachimiak / Alexander Leitner / Thomas Walzthoeni / Ruedi Aebersold / Wah Chiu / Judith Frydman /     Abstract: Maintaining proteostasis in eukaryotic protein folding involves cooperation of distinct chaperone systems. To understand how the essential ring-shaped chaperonin TRiC/CCT cooperates with the ...Maintaining proteostasis in eukaryotic protein folding involves cooperation of distinct chaperone systems. To understand how the essential ring-shaped chaperonin TRiC/CCT cooperates with the chaperone prefoldin/GIMc (PFD), we integrate cryoelectron microscopy (cryo-EM), crosslinking-mass-spectrometry and biochemical and cellular approaches to elucidate the structural and functional interplay between TRiC/CCT and PFD. We find these hetero-oligomeric chaperones associate in a defined architecture, through a conserved interface of electrostatic contacts that serves as a pivot point for a TRiC-PFD conformational cycle. PFD alternates between an open "latched" conformation and a closed "engaged" conformation that aligns the PFD-TRiC substrate binding chambers. PFD can act after TRiC bound its substrates to enhance the rate and yield of the folding reaction, suppressing non-productive reaction cycles. Disrupting the TRiC-PFD interaction in vivo is strongly deleterious, leading to accumulation of amyloid aggregates. The supra-chaperone assembly formed by PFD and TRiC is essential to prevent toxic conformations and ensure effective cellular proteostasis. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0492.map.gz emd_0492.map.gz | 49.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0492-v30.xml emd-0492-v30.xml emd-0492.xml emd-0492.xml | 29.5 KB 29.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0492_fsc.xml emd_0492_fsc.xml | 8.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_0492.png emd_0492.png | 55.3 KB | ||
| Filedesc metadata |  emd-0492.cif.gz emd-0492.cif.gz | 8.6 KB | ||
| Others |  emd_0492_additional.map.gz emd_0492_additional.map.gz emd_0492_half_map_1.map.gz emd_0492_half_map_1.map.gz emd_0492_half_map_2.map.gz emd_0492_half_map_2.map.gz | 40 MB 40.7 MB 40.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0492 http://ftp.pdbj.org/pub/emdb/structures/EMD-0492 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0492 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0492 | HTTPS FTP |
-Validation report
| Summary document |  emd_0492_validation.pdf.gz emd_0492_validation.pdf.gz | 829.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0492_full_validation.pdf.gz emd_0492_full_validation.pdf.gz | 828.9 KB | Display | |
| Data in XML |  emd_0492_validation.xml.gz emd_0492_validation.xml.gz | 14.2 KB | Display | |
| Data in CIF |  emd_0492_validation.cif.gz emd_0492_validation.cif.gz | 20.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0492 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0492 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0492 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0492 | HTTPS FTP |
-Related structure data
| Related structure data |  6nraMC  0490C  0491C  0493C  0494C  0495C  0496C  6nr8C  6nr9C  6nrbC  6nrcC  6nrdC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0492.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0492.map.gz / Format: CCP4 / Size: 52.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | hTRiC-hPFD Class1 (No PFD) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: hTRiC-hPFD Class1 (No PFD), unfiltered sum map
| File | emd_0492_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | hTRiC-hPFD Class1 (No PFD), unfiltered sum map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: hTRiC-hPFD Class1 (No PFD), half map 1
| File | emd_0492_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | hTRiC-hPFD Class1 (No PFD), half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: hTRiC-hPFD Class1 (No PFD), half map 2
| File | emd_0492_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | hTRiC-hPFD Class1 (No PFD), half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : hTRiC
| Entire | Name: hTRiC |
|---|---|
| Components |
|
-Supramolecule #1: hTRiC
| Supramolecule | Name: hTRiC / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: T-complex protein 1 subunit alpha
| Macromolecule | Name: T-complex protein 1 subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 58.061055 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: MEGPLSVFGD RSTGETIRSQ NVMAAASIAN IVKSSLGPVG LDKMLVDDIG DVTITNDGAT ILKLLEVEHP AAKVLCELAD LQDKEVGDG TTSVVIIAAE LLKNADELVK QKIHPTSVIS GYRLACKEAV RYINENLIVN TDELGRDCLI NAAKTSMSSK I IGINGDFF ...String: MEGPLSVFGD RSTGETIRSQ NVMAAASIAN IVKSSLGPVG LDKMLVDDIG DVTITNDGAT ILKLLEVEHP AAKVLCELAD LQDKEVGDG TTSVVIIAAE LLKNADELVK QKIHPTSVIS GYRLACKEAV RYINENLIVN TDELGRDCLI NAAKTSMSSK I IGINGDFF ANMVVDAVLA IKYTDIRGQP RYPVNSVNIL KAHGRSQMES MLISGYALNC VVGSQGMPKR IVNAKIACLD FS LQKTKMK LGVQVVITDP EKLDQIRQRE SDITKERIQK ILATGANVIL TTGGIDDMCL KYFVEAGAMA VRRVLKRDLK RIA KASGAT ILSTLANLEG EETFEAAMLG QAEEVVQERI CDDELILIKN TKARTSASII LRGANDFMCD EMERSLHDAL CVVK RVLES KSVVPGGGAV EAALSIYLEN YATSMGSREQ LAIAEFARSL LVIPNTLAVN AAQDSTDLVA KLRAFHNEAQ VNPER KNLK WIGLDLSNGK PRDNKQAGVF EPTIVKVKSL KFATEAAITI LRIDDLIKLH UniProtKB: T-complex protein 1 subunit alpha |
-Macromolecule #2: T-complex protein 1 subunit beta
| Macromolecule | Name: T-complex protein 1 subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.736742 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: ADEERAETAR LTSFIGAIAI GDLVKSTLGP KGMDKILLSS GRDASLMVTN DGATILKNIG VDNPAAKVLV DMSRVQDDEV GDGTTSVTV LAAELLREAE SLIAKKIHPQ TIIAGWREAT KAAREALLSS AVDHGSDEVK FRQDLMNIAG TTLSSKLLTH H KDHFTKLA ...String: ADEERAETAR LTSFIGAIAI GDLVKSTLGP KGMDKILLSS GRDASLMVTN DGATILKNIG VDNPAAKVLV DMSRVQDDEV GDGTTSVTV LAAELLREAE SLIAKKIHPQ TIIAGWREAT KAAREALLSS AVDHGSDEVK FRQDLMNIAG TTLSSKLLTH H KDHFTKLA VEAVLRLKGS GNLEAIHIIK KLGGSLADSY LDEGFLLDKK IGVNQPKRIE NAKILIANTG MDTDKIKIFG SR VRVDSTA KVAEIEHAEK EKMKEKVERI LKHGINCFIN RQLIYNYPEQ LFGAAGVMAI EHADFAGVER LALVTGGEIA STF DHPELV KLGSCKLIEE VMIGEDKLIH FSGVALGEAC TIVLRGATQQ ILDEAERSLH DALCVLAQTV KDSRTVYGGG CSEM LMAHA VTQLANRTPG KEAVAMESYA KALRMLPTII ADNAGYDSAD LVAQLRAAHS EGNTTAGLDM REGTIGDMAI LGITE SFQV KRQVLLSAAE AAEVILRVDN IIKAA UniProtKB: T-complex protein 1 subunit beta |
-Macromolecule #3: T-complex protein 1 subunit gamma
| Macromolecule | Name: T-complex protein 1 subunit gamma / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 57.165945 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: NTKRESGRKV QSGNINAAKT IADIIRTCLG PKSMMKMLLD PMGGIVMTND GNAILREIQV QHPAAKSMIE ISRTQDEEVG DGTTSVIIL AGEMLSVAEH FLEQQMHPTV VISAYRKALD DMISTLKKIS IPVDISDSDM MLNIINSSIT TKAISRWSSL A CNIALDAV ...String: NTKRESGRKV QSGNINAAKT IADIIRTCLG PKSMMKMLLD PMGGIVMTND GNAILREIQV QHPAAKSMIE ISRTQDEEVG DGTTSVIIL AGEMLSVAEH FLEQQMHPTV VISAYRKALD DMISTLKKIS IPVDISDSDM MLNIINSSIT TKAISRWSSL A CNIALDAV KMVQFEENGR KEIDIKKYAR VEKIPGGIIE DSCVLRGVMI NKDVTHPRMR RYIKNPRIVL LDSSLEYKKG ES QTDIEIT REEDFTRILQ MEEEYIQQLC EDIIQLKPDV VITEKGISDL AQHYLMRANI TAIRRVRKTD NNRIARACGA RIV SRPEEL REDDVGTGAG LLEIKKIGDE YFTFITDCKD PKACTILLRG ASKEILSEVE RNLQDAMQVC RNVLLDPQLV PGGG ASEMA VAHALTEKSK AMTGVEQWPY RAVAQALEVI PRTLIQNCGA STIRLLTSLR AKHTQENCET WGVNGETGTL VDMKE LGIW EPLAVKLQTY KTAVETAVLL LRIDDIVSG UniProtKB: T-complex protein 1 subunit gamma |
-Macromolecule #4: T-complex protein 1 subunit delta
| Macromolecule | Name: T-complex protein 1 subunit delta / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.6365 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: DRDKPAQIRF SNISAAKAVA DAIRTSLGPK GMDKMIQDGK GDVTITNDGA TILKQMQVLH PAARMLVELS KAQDIEAGDG TTSVVIIAG SLLDSCTKLL QKGIHPTIIS ESFQKALEKG IEILTDMSRP VELSDRETLL NSATTSLNSK VVSQYSSLLS P MSVNAVMK ...String: DRDKPAQIRF SNISAAKAVA DAIRTSLGPK GMDKMIQDGK GDVTITNDGA TILKQMQVLH PAARMLVELS KAQDIEAGDG TTSVVIIAG SLLDSCTKLL QKGIHPTIIS ESFQKALEKG IEILTDMSRP VELSDRETLL NSATTSLNSK VVSQYSSLLS P MSVNAVMK VIDPATATSV DLRDIKIVKK LGGTIDDCEL VEGLVLTQKV SNSGITRVEK AKIGLIQFCL SAPKTDMDNQ IV VSDYAQM DRVLREERAY ILNLVKQIKK TGCNVLLIQK SILRDALSDL ALHFLNKMKI MVIKDIERED IEFICKTIGT KPV AHIDQF TADMLGSAEL AEEVNLNGSG KLLKITGCAS PGKTVTIVVR GSNKLVIEEA ERSIHDALCV IRCLVKKRAL IAGG GAPEI ELALRLTEYS RTLSGMESYC VRAFADAMEV IPSTLAENAG LNPISTVTEL RNRHAQGEKT AGINVRKGGI SNILE ELVV QPLLVSVSAL TLATETVRSI LKIDDVVNTR UniProtKB: T-complex protein 1 subunit delta |
-Macromolecule #5: T-complex protein 1 subunit epsilon
| Macromolecule | Name: T-complex protein 1 subunit epsilon / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.989797 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: KSRLMGLEAL KSHIMAAKAV ANTMRTSLGP NGLDKMMVDK DGDVTVTNDG ATILSMMDVD HQIAKLMVEL SKSQDDEIGD GTTGVVVLA GALLEEAEQL LDRGIHPIRI ADGYEQAARV AIEHLDKISD SVLVDIKDTE PLIQTAKTTL GSKVVNSCHR Q MAEIAVNA ...String: KSRLMGLEAL KSHIMAAKAV ANTMRTSLGP NGLDKMMVDK DGDVTVTNDG ATILSMMDVD HQIAKLMVEL SKSQDDEIGD GTTGVVVLA GALLEEAEQL LDRGIHPIRI ADGYEQAARV AIEHLDKISD SVLVDIKDTE PLIQTAKTTL GSKVVNSCHR Q MAEIAVNA VLTVADMERR DVDFELIKVE GKVGGRLEDT KLIKGVIVDK DFSHPQMPKK VEDAKIAILT CPFEPPKPKT KH KLDVTSV EDYKALQKYE KEKFEEMIQQ IKETGANLAI CQWGFDDEAN HLLLQNNLPA VRWVGGPEIE LIAIATGGRI VPR FSELTA EKLGFAGLVQ EISFGTTKDK MLVIEQCKNS RAVTIFIRGG NKMIIEEAKR SLHDALCVIR NLIRDNRVVY GGGA AEISC ALAVSQEADK CPTLEQYAMR AFADALEVIP MALSENSGMN PIQTMTEVRA RQVKEMNPAL GIDCLHKGTN DMKQQ HVIE TLIGKKQQIS LATQMVRMIL KIDDIRKPGE SEE UniProtKB: T-complex protein 1 subunit epsilon |
-Macromolecule #6: T-complex protein 1 subunit zeta
| Macromolecule | Name: T-complex protein 1 subunit zeta / type: protein_or_peptide / ID: 6 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.445008 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: AEVARAQAAL AVNISAARGL QDVLRTNLGP KGTMKMLVSG AGDIKLTKDG NVLLHEMQIQ HPTASLIAKV ATAQDDITGD GTTSNVLII GELLKQADLY ISEGLHPRII TEGFEAAKEK ALQFLEEVKV SREMDRETLI DVARTSLRTK VHAELADVLT E AVVDSILA ...String: AEVARAQAAL AVNISAARGL QDVLRTNLGP KGTMKMLVSG AGDIKLTKDG NVLLHEMQIQ HPTASLIAKV ATAQDDITGD GTTSNVLII GELLKQADLY ISEGLHPRII TEGFEAAKEK ALQFLEEVKV SREMDRETLI DVARTSLRTK VHAELADVLT E AVVDSILA IKKQDEPIDL FMIEIMEMKH KSETDTSLIR GLVLDHGARH PDMKKRVEDA YILTCNVSLE YEKTEVNSGF FY KSAEERE KLVKAERKFI EDRVKKIIEL KRKVCGDSDK GFVVINQKGI DPFSLDALSK EGIVALRRAK RRNMERLTLA CGG VALNSF DDLSPDCLGH AGLVYEYTLG EEKFTFIEKC NNPRSVTLLI KGPNKHTLTQ IKDAVRDGLR AVKNAIDDGC VVPG AGAVE VAMAEALIKH KPSVKGRAQL GVQAFADALL IIPKVLAQNS GFDLQETLVK IQAEHSESGQ LVGVDLNTGE PMVAA EVGV WDNYCVKKQL LHSCTVIATN ILLVDEIMRA G UniProtKB: T-complex protein 1 subunit zeta |
-Macromolecule #7: T-complex protein 1 subunit eta
| Macromolecule | Name: T-complex protein 1 subunit eta / type: protein_or_peptide / ID: 7 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.369867 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: GTDSSQGIPQ LVSNISACQV IAEAVRTTLG PRGMDKLIVD GRGKATISND GATILKLLDV VHPAAKTLVD IAKSQDAEVG DGTTSVTLL AAEFLKQVKP YVEEGLHPQI IIRAFRTATQ LAVNKIKEIA VTVKKADKVE QRKLLEKCAM TALSSKLISQ Q KAFFAKMV ...String: GTDSSQGIPQ LVSNISACQV IAEAVRTTLG PRGMDKLIVD GRGKATISND GATILKLLDV VHPAAKTLVD IAKSQDAEVG DGTTSVTLL AAEFLKQVKP YVEEGLHPQI IIRAFRTATQ LAVNKIKEIA VTVKKADKVE QRKLLEKCAM TALSSKLISQ Q KAFFAKMV VDAVMMLDDL LQLKMIGIKK VQGGALEDSQ LVAGVAFKKT FSYAGFEMQP KKYHNPKIAL LNVELELKAE KD NAEIRVH TVEDYQAIVD AEWNILYDKL EKIHHSGAKV VLSKLPIGDV ATQYFADRDM FCAGRVPEED LKRTMMACGG SIQ TSVNAL SADVLGRCQV FEETQIGGER YNFFTGCPKA KTCTFILRGG AEQFMEETER SLHDAIMIVR RAIKNDSVVA GGGA IEMEL SKYLRDYSRT IPGKQQLLIG AYAKALEIIP RQLCDNAGFD ATNILNKLRA RHAQGGTWYG VDINNEDIAD NFEAF VWEP AMVRINALTA ASEAACLIVS VDETIKNPRS UniProtKB: T-complex protein 1 subunit eta |
-Macromolecule #8: T-complex protein 1 subunit theta
| Macromolecule | Name: T-complex protein 1 subunit theta / type: protein_or_peptide / ID: 8 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 56.102406 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: MLKEGAKHFS GLEEAVYRNI QACKELAQTT RTAYGPNGMN KMVINHLEKL FVTNDAATIL RELEVQHPAA KMIVMASHMQ EQEVGDGTN FVLVFAGALL ELAEELLRIG LSVSEVIEGY EIACRKAHEI LPNLVCCSAK NLRDIDEVSS LLRTSIMSKQ Y GNEVFLAK ...String: MLKEGAKHFS GLEEAVYRNI QACKELAQTT RTAYGPNGMN KMVINHLEKL FVTNDAATIL RELEVQHPAA KMIVMASHMQ EQEVGDGTN FVLVFAGALL ELAEELLRIG LSVSEVIEGY EIACRKAHEI LPNLVCCSAK NLRDIDEVSS LLRTSIMSKQ Y GNEVFLAK LIAQACVSIF PDSGHFNVDN IRVCKILGSG ISSSSVLHGM VFKKETEGDV TSVKDAKIAV YSCPFDGMIT ET KGTVLIK TAEELMNFSK GEENLMDAQV KAIADTGANV VVTGGKVADM ALHYANKYNI MLVRLNSKWD LRRLCKTVGA TAL PRLTPP VLEEMGHCDS VYLSEVGDTQ VVVFKHEKED GAISTIVLRG STDNLMDDIE RAVDDGVNTF KVLTRDKRLV PGGG ATEIE LAKQITSYGE TCPGLEQYAI KKFAEAFEAI PRALAENSGV KANEVISKLY AVHQEGNKNV GLDIEAEVPA VKDML EAGI LDTYLGKYWA IKLATNAAVT VLRVDQIIMA UniProtKB: T-complex protein 1 subunit theta |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 3200FSC |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)