[English] 日本語
 Yorodumi
Yorodumi- EMDB-25908: 2 microtubule protofilaments locally refined from Leucine Rich Re... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 2 microtubule protofilaments locally refined from Leucine Rich Repeat Kinase 2 filament structure bound to 11-protofilament microtubules | |||||||||
 Map data Map data | Sharpened map, recommend masking with provided refinement mask to remove partially resolved LRRK2 from the background. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | parkinson's disease / microtubule / CYTOSOLIC PROTEIN | |||||||||
| Biological species |  | |||||||||
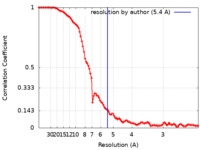
| Method | single particle reconstruction / cryo EM / Resolution: 5.4 Å | |||||||||
 Authors Authors | Matyszewski M / Leschziner AE | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2022 Journal: Nat Struct Mol Biol / Year: 2022Title: Structural basis for Parkinson's disease-linked LRRK2's binding to microtubules. Authors: David M Snead / Mariusz Matyszewski / Andrea M Dickey / Yu Xuan Lin / Andres E Leschziner / Samara L Reck-Peterson /  Abstract: Leucine-rich repeat kinase 2 (LRRK2) is one of the most commonly mutated genes in familial Parkinson's disease (PD). Under some circumstances, LRRK2 co-localizes with microtubules in cells, an ...Leucine-rich repeat kinase 2 (LRRK2) is one of the most commonly mutated genes in familial Parkinson's disease (PD). Under some circumstances, LRRK2 co-localizes with microtubules in cells, an association enhanced by PD mutations. We report a cryo-EM structure of the catalytic half of LRRK2, containing its kinase, in a closed conformation, and GTPase domains, bound to microtubules. We also report a structure of the catalytic half of LRRK1, which is closely related to LRRK2 but is not linked to PD. Although LRRK1's structure is similar to that of LRRK2, we find that LRRK1 does not interact with microtubules. Guided by these structures, we identify amino acids in LRRK2's GTPase that mediate microtubule binding; mutating them disrupts microtubule binding in vitro and in cells, without affecting LRRK2's kinase activity. Our results have implications for the design of therapeutic LRRK2 kinase inhibitors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25908.map.gz emd_25908.map.gz | 96.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25908-v30.xml emd-25908-v30.xml emd-25908.xml emd-25908.xml | 20.9 KB 20.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_25908_fsc.xml emd_25908_fsc.xml | 10.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_25908.png emd_25908.png | 81.7 KB | ||
| Masks |  emd_25908_msk_1.map emd_25908_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-25908.cif.gz emd-25908.cif.gz | 6 KB | ||
| Others |  emd_25908_additional_1.map.gz emd_25908_additional_1.map.gz emd_25908_half_map_1.map.gz emd_25908_half_map_1.map.gz emd_25908_half_map_2.map.gz emd_25908_half_map_2.map.gz | 50.6 MB 95.6 MB 95.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25908 http://ftp.pdbj.org/pub/emdb/structures/EMD-25908 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25908 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25908 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25908.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25908.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map, recommend masking with provided refinement mask to remove partially resolved LRRK2 from the background. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.16 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_25908_msk_1.map emd_25908_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Non-sharpened map
| File | emd_25908_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Non-sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 1
| File | emd_25908_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map 2
| File | emd_25908_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : LRRK2RCKW filament bound to a 11-pf microtubule with MLi-2 presen...
| Entire | Name: LRRK2RCKW filament bound to a 11-pf microtubule with MLi-2 present. This map focuses on the microtubule only |
|---|---|
| Components |
|
-Supramolecule #1: LRRK2RCKW filament bound to a 11-pf microtubule with MLi-2 presen...
| Supramolecule | Name: LRRK2RCKW filament bound to a 11-pf microtubule with MLi-2 present. This map focuses on the microtubule only type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Macromolecule #1: Tubulin alpha
| Macromolecule | Name: Tubulin alpha / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDSFN TFFSETGAGK HVPRAVFVD LEPTVIDEVR TGTYRQLFHP EQLITGKEDA ANNYARGHYT IGKEIIDLVL D RIRKLADQ CTGLQGFLVF HSFGGGTGSG FTSLLMERLS VDYGKKSKLE ...String: MRECISIHVG QAGVQIGNAC WELYCLEHGI QPDGQMPSDK TIGGGDDSFN TFFSETGAGK HVPRAVFVD LEPTVIDEVR TGTYRQLFHP EQLITGKEDA ANNYARGHYT IGKEIIDLVL D RIRKLADQ CTGLQGFLVF HSFGGGTGSG FTSLLMERLS VDYGKKSKLE FSIYPAPQVS TA VVEPYNS ILTTHTTLEH SDCAFMVDNE AIYDICRRNL DIERPTYTNL NRLIGQIVSS ITA SLRFDG ALNVDLTEFQ TNLVPYPRIH FPLATYAPVI SAEKAYHEQL SVAEITNACF EPAN QMVKC DPRHGKYMAC CLLYRGDVVP KDVNAAIATI KTKRTIQFVD WCPTGFKVGI NYQPP TVVP GGDLAKVQRA VCMLSNTTAI AEAWARLDHK FDLMYAKRAF VHWYVGEGME EGEFSE ARE DMAALEKDYE EVGVDSVEGE GEEEGEEY |
-Macromolecule #2: Tubulin beta
| Macromolecule | Name: Tubulin beta / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Sequence | String: MREIVHIQIG QCGNQIGAKF WEVIGEEHGI DWAGSYCGDS ALQLERISVY YNEAHGKKYV PRAVLVDLE PGTMDSIRSS RVGALFQPDS FVHGNSGAGN NWAKGYYTEG AELVDRVLDA V RTEAEGCD CLQGFQLVHS LGGGTGSGMG TLLLGKIREE YPDRILNSFS ...String: MREIVHIQIG QCGNQIGAKF WEVIGEEHGI DWAGSYCGDS ALQLERISVY YNEAHGKKYV PRAVLVDLE PGTMDSIRSS RVGALFQPDS FVHGNSGAGN NWAKGYYTEG AELVDRVLDA V RTEAEGCD CLQGFQLVHS LGGGTGSGMG TLLLGKIREE YPDRILNSFS VMPSPKVSDT VV EPYNAVL ALHQLVLNSD ACFCIDNEAL YDICFRTLRL STPTYGDLNH LVSLTMSGIT TSL RFPGQL NADLRKLAVN MVPFPRLHFF MPGFAPLTAQ GSQQYRALTV AELTQQMFDA RNTM AACDP RRGRYLTVAC IFRGRMSTKE VDEQLLNVQT RNSSCFVEWI PNNVKVAVCD IPPRG LSMA ATFIGNNTAI QELFSRISEH FSAMFKRKAF VHWYTGEGMD INEFTEAESN IQDLVS EYQ QFQDARADVE EEEIGGEAEV EPADKEH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
Details: This is the final dilution buffer. The incubation buffer consisted of 1x BRB80, 10% glycerol, 1mM DTT, 1mM GTP, 1mM MgCl2, 10 uM taxol, and 5 uM MLi-2. Sample was diluted 3-fold right before ...Details: This is the final dilution buffer. The incubation buffer consisted of 1x BRB80, 10% glycerol, 1mM DTT, 1mM GTP, 1mM MgCl2, 10 uM taxol, and 5 uM MLi-2. Sample was diluted 3-fold right before freezing with the final buffer. | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Homemade / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: LACEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 45 sec. / Details: Using EMS LC-300 lacey grids (Not homemade) | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K | ||||||||||||||||||
| Details | 4.5 uM of LRRK2RCKW was allowed to incubate with 2.25 uM of tubulin dimer, causing both to co-polymerize. 5 uM of MLi-2 was present as well. The sample was diluted 3-fold right before freezing (1.5 uM LRRK2RCKW concentration final). |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Average exposure time: 10.0 sec. / Average electron dose: 55.0 e/Å2 / Details: 250 ms frames |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 36000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)