[English] 日本語
 Yorodumi
Yorodumi- EMDB-13845: Protein community member pyruvate dehydrogenase complex E2 core f... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13845 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Protein community member pyruvate dehydrogenase complex E2 core from C. thermophilum | |||||||||
 Map data Map data | C. thermophilum Native Community Member Pyruvate Dehydrogenase Complex E2 Core | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Dihydrolipoyl / transacetylase / E2 / Pyruvate / TRANSFERASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationdihydrolipoyllysine-residue acetyltransferase / dihydrolipoyllysine-residue acetyltransferase activity / pyruvate decarboxylation to acetyl-CoA / pyruvate dehydrogenase complex / mitochondrion Similarity search - Function | |||||||||
| Biological species |  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) / Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) /  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) | |||||||||
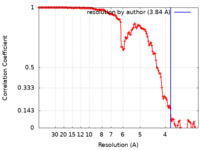
| Method | single particle reconstruction / cryo EM / Resolution: 3.84 Å | |||||||||
 Authors Authors | Skalidis I / Kyrilis FL / Tueting C / Hamdi F / Kastritis PL | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Cryo-EM and artificial intelligence visualize endogenous protein community members. Authors: Ioannis Skalidis / Fotis L Kyrilis / Christian Tüting / Farzad Hamdi / Grzegorz Chojnowski / Panagiotis L Kastritis /  Abstract: Cellular function is underlined by megadalton assemblies organizing in proximity, forming communities. Metabolons are protein communities involving metabolic pathways such as protein, fatty acid, and ...Cellular function is underlined by megadalton assemblies organizing in proximity, forming communities. Metabolons are protein communities involving metabolic pathways such as protein, fatty acid, and thioesters of coenzyme-A synthesis. Metabolons are highly heterogeneous due to their function, making their analysis particularly challenging. Here, we simultaneously characterize metabolon-embedded architectures of a 60S pre-ribosome, fatty acid synthase, and pyruvate/oxoglutarate dehydrogenase complex E2 cores de novo. Cryo-electron microscopy (cryo-EM) 3D reconstructions are resolved at 3.84-4.52 Å resolution by collecting <3,000 micrographs of a single cellular fraction. After combining cryo-EM with artificial intelligence-based atomic modeling and de novo sequence identification methods, at this resolution range, polypeptide hydrogen bonding patterns are discernible. Residing molecular components resemble their purified counterparts from other eukaryotes but also exhibit substantial conformational variation with potential functional implications. Our results propose an integrated tool, boosted by machine learning, that opens doors for structural systems biology spearheaded by cryo-EM characterization of native cell extracts. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13845.map.gz emd_13845.map.gz | 204 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13845-v30.xml emd-13845-v30.xml emd-13845.xml emd-13845.xml | 19.6 KB 19.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13845_fsc.xml emd_13845_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_13845.png emd_13845.png | 82.8 KB | ||
| Masks |  emd_13845_msk_1.map emd_13845_msk_1.map | 216 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13845.cif.gz emd-13845.cif.gz | 5.9 KB | ||
| Others |  emd_13845_additional_1.map.gz emd_13845_additional_1.map.gz emd_13845_half_map_1.map.gz emd_13845_half_map_1.map.gz emd_13845_half_map_2.map.gz emd_13845_half_map_2.map.gz | 51.3 MB 200.4 MB 200.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13845 http://ftp.pdbj.org/pub/emdb/structures/EMD-13845 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13845 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13845 | HTTPS FTP |
-Related structure data
| Related structure data |  7q5rMC  7q5qC  7q5sC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10892 (Title: Cryo-EM SPA dataset of Megadalton-range protein communities from a Chaetomium thermophilum native cell extract EMPIAR-10892 (Title: Cryo-EM SPA dataset of Megadalton-range protein communities from a Chaetomium thermophilum native cell extractData size: 1.1 TB Data #1: Unaligned fractions saved by Falcon 3 EC camera [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13845.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13845.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C. thermophilum Native Community Member Pyruvate Dehydrogenase Complex E2 Core | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.5678 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_13845_msk_1.map emd_13845_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: C. thermophilum Native Community Member Signature 2 Ab Initio Model
| File | emd_13845_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C. thermophilum Native Community Member Signature 2 Ab Initio Model | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: C. thermophilum Native Community Member Pyruvate Dehydrogenase Complex...
| File | emd_13845_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C. thermophilum Native Community Member Pyruvate Dehydrogenase Complex E2 Core, Half-Map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: C. thermophilum Native Community Member Pyruvate Dehydrogenase Complex...
| File | emd_13845_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | C. thermophilum Native Community Member Pyruvate Dehydrogenase Complex E2 Core, Half-Map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Native 60-mer core of Pyruvate Dehydrogenase Complex
| Entire | Name: Native 60-mer core of Pyruvate Dehydrogenase Complex |
|---|---|
| Components |
|
-Supramolecule #1: Native 60-mer core of Pyruvate Dehydrogenase Complex
| Supramolecule | Name: Native 60-mer core of Pyruvate Dehydrogenase Complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) |
-Macromolecule #1: Acetyltransferase component of pyruvate dehydrogenase complex
| Macromolecule | Name: Acetyltransferase component of pyruvate dehydrogenase complex type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO / EC number: dihydrolipoyllysine-residue acetyltransferase |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 48.777488 KDa |
| Sequence | String: MLAQVLRRQA LQHVRLARAA APSLTRWYAS YPPHTIVKMP ALSPTMTSGN IGAWQKKPGD AITPGEVLVE IETDKAQMDF EFQEEGVLA KILKETGEKD VAVGSPIAVL VEEGTDINAF QNFTLEDAGG DAAAPAAPAK EELAKAETAP TPASTSAPEP E ETTSTGKL ...String: MLAQVLRRQA LQHVRLARAA APSLTRWYAS YPPHTIVKMP ALSPTMTSGN IGAWQKKPGD AITPGEVLVE IETDKAQMDF EFQEEGVLA KILKETGEKD VAVGSPIAVL VEEGTDINAF QNFTLEDAGG DAAAPAAPAK EELAKAETAP TPASTSAPEP E ETTSTGKL EPALDREPNV SFAAKKLAHE LDVPLKALKG TGPGGKITEE DVKKAASAPA AAAAAPGAAY QDIPISNMRK TI ATRLKES VSENPHFFVT SELSVSKLLK LRQALNSSAE GRYKLSVNDF LIKAIAVACK RVPAVNSSWR DGVIRQFDTV DVS VAVATP TGLITPIVKG VEAKGLETIS ATVKELAKKA RDGKLKPEDY QGGTISISNM GMNPAVERFT AIINPPQAAI LAVG TTKKV AVPVENEDGT TGVEWDDQIV VTASFDHKVV DGAVGAEWMR ELKKVVENPL ELLL UniProtKB: Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Component - Concentration: 200.0 mM / Component - Formula: NH4CH2COOH / Component - Name: Ammonium acetate |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: For plunging, blot force 0 and blotting time of 4 sec were applied.. |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Temperature | Min: 77.15 K / Max: 103.15 K |
| Alignment procedure | Coma free - Residual tilt: 14.7 mrad |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Number real images: 2808 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 92000 |
| Sample stage | Specimen holder model: OTHER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Initial models were predicted using AlphaFold v2.0.1, fitted into reconstructions using COOT and finally refined in real space using COOT and phenix.real_space_refine |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-7q5r: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)