[English] 日本語
 Yorodumi
Yorodumi- EMDB-10835: High resolution cryo-EM structure of urease from the pathogen Yer... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10835 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | High resolution cryo-EM structure of urease from the pathogen Yersinia enterocolitica | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Urease / enzyme / nickel / metalloenzyme / pathogen / METAL BINDING PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationurease complex / urease / urease activity / urea catabolic process / nickel cation binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Yersinia enterocolitica W22703 (bacteria) Yersinia enterocolitica W22703 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 1.98 Å | |||||||||
 Authors Authors | Righetto RD / Anton L | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: High-resolution cryo-EM structure of urease from the pathogen Yersinia enterocolitica. Authors: Ricardo D Righetto / Leonie Anton / Ricardo Adaixo / Roman P Jakob / Jasenko Zivanov / Mohamed-Ali Mahi / Philippe Ringler / Torsten Schwede / Timm Maier / Henning Stahlberg /  Abstract: Urease converts urea into ammonia and carbon dioxide and makes urea available as a nitrogen source for all forms of life except animals. In human bacterial pathogens, ureases also aid in the invasion ...Urease converts urea into ammonia and carbon dioxide and makes urea available as a nitrogen source for all forms of life except animals. In human bacterial pathogens, ureases also aid in the invasion of acidic environments such as the stomach by raising the surrounding pH. Here, we report the structure of urease from the pathogen Yersinia enterocolitica at 2 Å resolution from cryo-electron microscopy. Y. enterocolitica urease is a dodecameric assembly of a trimer of three protein chains, ureA, ureB and ureC. The high data quality enables detailed visualization of the urease bimetal active site and of the impact of radiation damage. The obtained structure is of sufficient quality to support drug development efforts. #1:  Journal: Biorxiv / Year: 2020 Journal: Biorxiv / Year: 2020Title: High-resolution cryo-EM structure of urease from the pathogen Yersinia enterocolitica Authors: Righetto RD / Anton L / Adaixo R / Jakob R / Zivanov J / Mahi MA / Ringler P / Schwede T / Maier T / Stahlberg H | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10835.map.gz emd_10835.map.gz | 302.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10835-v30.xml emd-10835-v30.xml emd-10835.xml emd-10835.xml | 23.4 KB 23.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10835_fsc.xml emd_10835_fsc.xml | 18 KB | Display |  FSC data file FSC data file |
| Images |  emd_10835.png emd_10835.png | 169.2 KB | ||
| Masks |  emd_10835_msk_1.map emd_10835_msk_1.map | 512 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10835.cif.gz emd-10835.cif.gz | 7 KB | ||
| Others |  emd_10835_half_map_1.map.gz emd_10835_half_map_1.map.gz emd_10835_half_map_2.map.gz emd_10835_half_map_2.map.gz | 410.4 MB 410.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10835 http://ftp.pdbj.org/pub/emdb/structures/EMD-10835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10835 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10835 | HTTPS FTP |
-Related structure data
| Related structure data |  6yl3MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10389 (Title: High resolution cryo-EM structure of urease from the pathogen Yersinia enterocolitica EMPIAR-10389 (Title: High resolution cryo-EM structure of urease from the pathogen Yersinia enterocoliticaData size: 839.9 Data #1: Aligned averages of Y. enterocolitica urease [micrographs - single frame] Data #2: Raw multi-frame micrographs of Y. enterocolitica urease [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10835.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10835.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.639 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10835_msk_1.map emd_10835_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
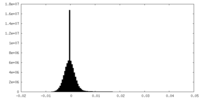
| Density Histograms |
-Half map: #1
| File | emd_10835_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_10835_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Urease oligomer from Y. enterocolitica
| Entire | Name: Urease oligomer from Y. enterocolitica |
|---|---|
| Components |
|
-Supramolecule #1: Urease oligomer from Y. enterocolitica
| Supramolecule | Name: Urease oligomer from Y. enterocolitica / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Yersinia enterocolitica W22703 (bacteria) / Strain: E40 Yersinia enterocolitica W22703 (bacteria) / Strain: E40 |
| Molecular weight | Theoretical: 1.025 MDa |
-Macromolecule #1: Urease subunit gamma
| Macromolecule | Name: Urease subunit gamma / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: urease |
|---|---|
| Source (natural) | Organism:  Yersinia enterocolitica W22703 (bacteria) Yersinia enterocolitica W22703 (bacteria) |
| Molecular weight | Theoretical: 11.063837 KDa |
| Sequence | String: MQLTPREVEK LMIYTLSDVA FKRKARGLKL NYPEAVSIIT VTAMEGARDG KSVEDVMKEA SKVLTKDDVM DGVADLIPNV QVEAIFTDG SRLVTVHDPI K UniProtKB: Urease subunit gamma |
-Macromolecule #2: Urease subunit beta
| Macromolecule | Name: Urease subunit beta / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO / EC number: urease |
|---|---|
| Source (natural) | Organism:  Yersinia enterocolitica W22703 (bacteria) Yersinia enterocolitica W22703 (bacteria) |
| Molecular weight | Theoretical: 14.611317 KDa |
| Sequence | String: SEQNTPLGGC ILADTPITFN ENKPVTKVKV RNTGDRPIQV GSHFHFFEVN RALEFDRAAA YGKRLNISST TAIRFEPGDE TEVPLIPFG GKQTLYGFNN LVDGWTGEGV VPNSERPDKL EAIRRAAERG FKS UniProtKB: Urease subunit beta |
-Macromolecule #3: Urease subunit alpha
| Macromolecule | Name: Urease subunit alpha / type: protein_or_peptide / ID: 3 / Number of copies: 12 / Enantiomer: LEVO / EC number: urease |
|---|---|
| Source (natural) | Organism:  Yersinia enterocolitica W22703 (bacteria) Yersinia enterocolitica W22703 (bacteria) |
| Molecular weight | Theoretical: 61.054859 KDa |
| Sequence | String: PQISRQEYAG LFGPTTGDKI RLGDTNLFIE IEKDLRGYGE ESVYGGGKSL RDGMGANNHL TRDNGVLDLV ITNVTIVDAR LGVIKADVG IRDGKIAGIG KSGNPGVMDG VTPGLVVGVS TDAISGEHLI LTAAGIDTHI HLISPQQAYH ALSNGVATFF G GGIGPTDG ...String: PQISRQEYAG LFGPTTGDKI RLGDTNLFIE IEKDLRGYGE ESVYGGGKSL RDGMGANNHL TRDNGVLDLV ITNVTIVDAR LGVIKADVG IRDGKIAGIG KSGNPGVMDG VTPGLVVGVS TDAISGEHLI LTAAGIDTHI HLISPQQAYH ALSNGVATFF G GGIGPTDG TNGTTVTPGP WNIRQMLRSV EGLPVNVGIL GKGNSYGRGP LLEQAIAGVV GY(KCX)VHEDWGA TANALRHS L RMADEMDIQV SVHTDSLNEC GYVEDTIDAF EGRTIHTFHT EGAGGGHAPD IIRVASQPNV LPSSTNPTLP YGVNSQAEL FDMIMVCHNL NPNVPADVSF AESRVRPETI AAENVLHDMG VISMFSSDSQ AMGRVGENWL RVMQTANAMK ASRGKLPEDA PGNDNFRVL RYVAKITINP AIAQGVSHVI GSVEVGKMAD LVLWDPRFFG AKPKMVIKGG MINWAAMGDP NASLPTPQPV F YRPMFGAM GKTMQDTCVT FVSQAALDDG VKEKAGLDRQ VIAVKNCRTI SKHDLVRNDQ TPNIEVDPET FAVKVDGVHA TC EPIDTAA MNQRYFFG UniProtKB: Urease subunit alpha |
-Macromolecule #4: NICKEL (II) ION
| Macromolecule | Name: NICKEL (II) ION / type: ligand / ID: 4 / Number of copies: 24 / Formula: NI |
|---|---|
| Molecular weight | Theoretical: 58.693 Da |
| Chemical component information |  ChemComp-NI: |
-Macromolecule #5: water
| Macromolecule | Name: water / type: ligand / ID: 5 / Number of copies: 3672 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.39 mg/mL |
|---|---|
| Buffer | pH: 7 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293.15 K / Instrument: FEI VITROBOT MARK IV / Details: 3 seconds blotting. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)