+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0717 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Troponin of cardiac thin filament in low-calcium state | |||||||||
 Map data Map data | Cardiac troponin in low-calcium state | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cardiac thin filament / CONTRACTILE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationatrial cardiac muscle tissue morphogenesis / regulation of systemic arterial blood pressure by ischemic conditions / troponin C binding / Striated Muscle Contraction / contractile muscle fiber / diaphragm contraction / regulation of muscle filament sliding speed / troponin T binding / cardiac Troponin complex / cardiac myofibril ...atrial cardiac muscle tissue morphogenesis / regulation of systemic arterial blood pressure by ischemic conditions / troponin C binding / Striated Muscle Contraction / contractile muscle fiber / diaphragm contraction / regulation of muscle filament sliding speed / troponin T binding / cardiac Troponin complex / cardiac myofibril / actin crosslink formation / negative regulation of ATP-dependent activity / troponin complex / regulation of muscle contraction / regulation of smooth muscle contraction / positive regulation of ATP-dependent activity / transition between fast and slow fiber / Ion homeostasis / muscle filament sliding / regulation of cardiac muscle contraction by calcium ion signaling / response to metal ion / cardiac muscle cell contraction / sarcomere organization / ventricular cardiac muscle tissue morphogenesis / heart contraction / tropomyosin binding / regulation of heart contraction / troponin I binding / myofibril / striated muscle thin filament / vasculogenesis / calcium channel inhibitor activity / striated muscle contraction / skeletal muscle contraction / cardiac muscle contraction / muscle contraction / sarcomere / response to bacterium / response to calcium ion / structural constituent of cytoskeleton / intracellular calcium ion homeostasis / calcium-dependent protein binding / actin filament binding / heart development / actin binding / protein-macromolecule adaptor activity / protein domain specific binding / calcium ion binding / protein kinase binding / protein-containing complex binding / protein homodimerization activity / identical protein binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
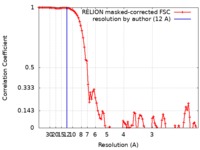
| Method | single particle reconstruction / cryo EM / Resolution: 12.0 Å | |||||||||
 Authors Authors | Oda T / Yanagisawa H | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2020 Journal: J Struct Biol / Year: 2020Title: Cryo-EM structures of cardiac thin filaments reveal the 3D architecture of troponin. Authors: Toshiyuki Oda / Haruaki Yanagisawa / Takeyuki Wakabayashi /  Abstract: Troponin is an essential component of striated muscle and it regulates the sliding of actomyosin system in a calcium-dependent manner. Despite its importance, the structure of troponin has been ...Troponin is an essential component of striated muscle and it regulates the sliding of actomyosin system in a calcium-dependent manner. Despite its importance, the structure of troponin has been elusive due to its high structural heterogeneity. In this study, we analyzed the 3D structures of murine cardiac thin filaments using a cryo-electron microscope equipped with a Volta phase plate (VPP). Contrast enhancement by a VPP enabled us to reconstruct the entire repeat of the thin filament. We determined the orientation of troponin relative to F-actin and tropomyosin, and characterized the interactions between troponin and tropomyosin. This study provides a structural basis for understanding the molecular mechanism of actomyosin system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0717.map.gz emd_0717.map.gz | 43.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0717-v30.xml emd-0717-v30.xml emd-0717.xml emd-0717.xml | 13.9 KB 13.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0717_fsc.xml emd_0717_fsc.xml | 9.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_0717.png emd_0717.png | 71.8 KB | ||
| Filedesc metadata |  emd-0717.cif.gz emd-0717.cif.gz | 5.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0717 http://ftp.pdbj.org/pub/emdb/structures/EMD-0717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0717 | HTTPS FTP |
-Related structure data
| Related structure data |  6kltMC  0711C  0712C  0714C  0715C  0718C  0796C  0797C  0798C  0799C  0802C  0804C  0805C  0806C  0807C  0808C  6kllC  6klnC  6klpC  6klqC  6kluC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10348 (Title: Cardiac thin filament in low calcium state / Data size: 2.5 TB EMPIAR-10348 (Title: Cardiac thin filament in low calcium state / Data size: 2.5 TBData #1: Unaligned multiframe micrographs of cardiac myofilaments in low calcium state [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0717.map.gz / Format: CCP4 / Size: 70.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0717.map.gz / Format: CCP4 / Size: 70.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cardiac troponin in low-calcium state | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : F-actin of cardiac thin filament
| Entire | Name: F-actin of cardiac thin filament |
|---|---|
| Components |
|
-Supramolecule #1: F-actin of cardiac thin filament
| Supramolecule | Name: F-actin of cardiac thin filament / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Troponin C, slow skeletal and cardiac muscles
| Macromolecule | Name: Troponin C, slow skeletal and cardiac muscles / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 16.644533 KDa |
| Sequence | String: TEEQKNEFKA AFDIFVLGAD GCISTKELGK VMRMLGQNPT PEELQEMIDE VDEDGSGTVD FDEFLVMMVR CMKDDSKGKS EEELSDLFR MFDKNADGYI DLDELKMMLQ ATGETITEDD IEELMKDGDK NNDGRIDYDE FLEFMK UniProtKB: Troponin C, slow skeletal and cardiac muscles |
-Macromolecule #2: Troponin T, cardiac muscle
| Macromolecule | Name: Troponin T, cardiac muscle / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.735045 KDa |
| Sequence | String: GKRQTEREKK KKILAERRKA LAIDHLNEDQ LREKAKELWQ SIHNLEAEKF DLQEKFKQQK YEINVLRNRI N UniProtKB: Troponin T, cardiac muscle |
-Macromolecule #3: Troponin I, cardiac muscle
| Macromolecule | Name: Troponin I, cardiac muscle / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.524673 KDa |
| Sequence | String: QLKTLMLQIA KQEMEREAEE RRGEKGRVLR TRCQPLELDG LGFEELQDLC RQLHARVDKV DEERYDVEAK VTKNITEIAD LTQKIYDLR GKFKRPTLRR VRISADAMMQ ALLGTR UniProtKB: Troponin I, cardiac muscle |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7.2 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE / Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average exposure time: 5.6 sec. / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.1 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)