+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6903 | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ebola virus nucleoprotein-RNA complex | ||||||||||||||||||||||||||||||
 Map data Map data | Ebolavirus nucleoprotein RNA complex (biological assembly) | ||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||
 Keywords Keywords | ebolavirus / RNA / nucleoprotein / nucleocapsid / helical / VIRAL PROTEIN | ||||||||||||||||||||||||||||||
| Function / homology | Ebola nucleoprotein / Ebola nucleoprotein / viral RNA genome packaging / helical viral capsid / viral nucleocapsid / host cell cytoplasm / ribonucleoprotein complex / RNA binding / Nucleoprotein Function and homology information Function and homology information | ||||||||||||||||||||||||||||||
| Biological species |   Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | ||||||||||||||||||||||||||||||
| Method | helical reconstruction / cryo EM / Resolution: 3.6 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Sugita Y / Matsunami H / Kawaoka Y / Noda T / Wolf M | ||||||||||||||||||||||||||||||
| Funding support |  Japan, 9 items Japan, 9 items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2018 Journal: Nature / Year: 2018Title: Cryo-EM structure of the Ebola virus nucleoprotein-RNA complex at 3.6 Å resolution. Authors: Yukihiko Sugita / Hideyuki Matsunami / Yoshihiro Kawaoka / Takeshi Noda / Matthias Wolf /   Abstract: Ebola virus causes haemorrhagic fever with a high fatality rate in humans and non-human primates. It belongs to the family Filoviridae in the order Mononegavirales, which are viruses that contain ...Ebola virus causes haemorrhagic fever with a high fatality rate in humans and non-human primates. It belongs to the family Filoviridae in the order Mononegavirales, which are viruses that contain linear, non-segmented, negative-sense, single-stranded genomic RNA. The enveloped, filamentous virion contains the nucleocapsid, consisting of the helical nucleoprotein-RNA complex, VP24, VP30, VP35 and viral polymerase. The nucleoprotein-RNA complex acts as a scaffold for nucleocapsid formation and as a template for RNA replication and transcription by condensing RNA into the virion. RNA binding and nucleoprotein oligomerization are synergistic and do not readily occur independently. Although recent cryo-electron tomography studies have revealed the overall architecture of the nucleocapsid core, there has been no high-resolution reconstruction of the nucleocapsid. Here we report the structure of a recombinant Ebola virus nucleoprotein-RNA complex expressed in mammalian cells without chemical fixation, at near-atomic resolution using single-particle cryo-electron microscopy. Our structure reveals how the Ebola virus nucleocapsid core encapsidates its viral genome, its sequence-independent coordination with RNA by nucleoprotein, and the dynamic transition between the RNA-free and RNA-bound states. It provides direct structural evidence for the role of the N terminus of nucleoprotein in subunit oligomerization, and for the hydrophobic and electrostatic interactions that lead to the formation of the helical assembly. The structure is validated as representative of the native biological assembly of the nucleocapsid core by consistent dimensions and symmetry with the full virion. The atomic model provides a detailed mechanistic basis for understanding nucleocapsid assembly and highlights key structural features that may serve as targets for anti-viral drug development. | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6903.map.gz emd_6903.map.gz | 59.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6903-v30.xml emd-6903-v30.xml emd-6903.xml emd-6903.xml | 17.4 KB 17.4 KB | Display Display |  EMDB header EMDB header |
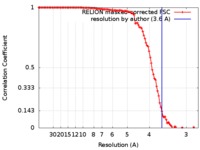
| FSC (resolution estimation) |  emd_6903_fsc.xml emd_6903_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_6903.png emd_6903.png | 238 KB | ||
| Filedesc metadata |  emd-6903.cif.gz emd-6903.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6903 http://ftp.pdbj.org/pub/emdb/structures/EMD-6903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6903 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6903 | HTTPS FTP |
-Related structure data
| Related structure data |  5z9wMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6903.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6903.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ebolavirus nucleoprotein RNA complex (biological assembly) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.39 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ebolavirus nucleoprotein RNA complex (biological assembly)
| Entire | Name: Ebolavirus nucleoprotein RNA complex (biological assembly) |
|---|---|
| Components |
|
-Supramolecule #1: Ebolavirus nucleoprotein RNA complex (biological assembly)
| Supramolecule | Name: Ebolavirus nucleoprotein RNA complex (biological assembly) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 166 KDa |
-Macromolecule #1: Ebolavirus nucleoprotein (residues 19-406)
| Macromolecule | Name: Ebolavirus nucleoprotein (residues 19-406) / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.496086 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MDYHKILTAG LSVQQGIVRQ RVIPVYQVNN LEEICQLIIQ AFEAGVDFQE SADSFLLMLC LHHAYQGDYK LFLESGAVKY LEGHGFRFE VKKRDGVKRL EELLPAVSSG KNIKRTLAAM PEEETTEANA GQFLSFASLF LPKLVVGEKA CLEKVQRQIQ V HAEQGLIQ ...String: MDYHKILTAG LSVQQGIVRQ RVIPVYQVNN LEEICQLIIQ AFEAGVDFQE SADSFLLMLC LHHAYQGDYK LFLESGAVKY LEGHGFRFE VKKRDGVKRL EELLPAVSSG KNIKRTLAAM PEEETTEANA GQFLSFASLF LPKLVVGEKA CLEKVQRQIQ V HAEQGLIQ YPTAWQSVGH MMVIFRLMRT NFLIKFLLIH QGMHMVAGHD ANDAVISNSV AQARFSGLLI VKTVLDHILQ KT ERGVRLH PLARTAKVKN EVNSFKAALS SLAKHGEYAP FARLLNLSGV NNLEHGLFPQ LSAIALGVAT AHGSTLAGVN VGE QYQQLR EAATEAEKQL QQYAESRELD HLGLDDQEKK ILMNFHQKKN EISFQQTNAM VTLRKERLAK LT |
-Macromolecule #2: RNA (6-MER)
| Macromolecule | Name: RNA (6-MER) / type: rna / ID: 2 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 1.792037 KDa |
| Sequence | String: UUUUUU |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | helical array |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 Component:
| ||||||||||||
| Grid | Model: C-flat-1.2/1.3 4C / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 150 / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 20 sec. / Pretreatment - Atmosphere: OTHER / Details: Gatan Solarus | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 289 K / Instrument: FEI VITROBOT MARK IV / Details: 3 second blot, 2.5uL. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77.0 K / Max: 100.0 K |
| Alignment procedure | Coma free - Residual tilt: 0.1 mrad |
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Details | nanoprobe, parallel beam illumination |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 7676 pixel / Digitization - Dimensions - Height: 7420 pixel / Digitization - Frames/image: 1-75 / Number grids imaged: 1 / Number real images: 2467 / Average exposure time: 15.0 sec. / Average electron dose: 105.0 e/Å2 Details: frame alignment and dose weighting using motioncor2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 4.0 µm / Calibrated defocus min: 0.8 µm / Calibrated magnification: 47619 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-5z9w: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)