+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6455 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | TRPV2 ion channel from rabbit | |||||||||
 Map data Map data | TRPV2, sharpened with a B factor of -76 using RELION postprocess | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRP channel / ion transport | |||||||||
| Function / homology |  Function and homology information Function and homology informationgrowth cone membrane / response to temperature stimulus / positive regulation of calcium ion import / calcium ion import across plasma membrane / positive regulation of axon extension / axonal growth cone / calcium channel activity / positive regulation of cold-induced thermogenesis / cell body / cell surface / identical protein binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Herzik MA / Zubcevic L / Chung BC / Lander GC / Lee SY | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2016 Journal: Nat Struct Mol Biol / Year: 2016Title: Cryo-electron microscopy structure of the TRPV2 ion channel. Authors: Lejla Zubcevic / Mark A Herzik / Ben C Chung / Zhirui Liu / Gabriel C Lander / Seok-Yong Lee /  Abstract: Transient receptor potential vanilloid (TRPV) cation channels are polymodal sensors involved in a variety of physiological processes. TRPV2, a member of the TRPV family, is regulated by temperature, ...Transient receptor potential vanilloid (TRPV) cation channels are polymodal sensors involved in a variety of physiological processes. TRPV2, a member of the TRPV family, is regulated by temperature, by ligands, such as probenecid and cannabinoids, and by lipids. TRPV2 has been implicated in many biological functions, including somatosensation, osmosensation and innate immunity. Here we present the atomic model of rabbit TRPV2 in its putative desensitized state, as determined by cryo-EM at a nominal resolution of ∼4 Å. In the TRPV2 structure, the transmembrane segment 6 (S6), which is involved in gate opening, adopts a conformation different from the one observed in TRPV1. Structural comparisons of TRPV1 and TRPV2 indicate that a rotation of the ankyrin-repeat domain is coupled to pore opening via the TRP domain, and this pore opening can be modulated by rearrangements in the secondary structure of S6. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6455.map.gz emd_6455.map.gz | 14.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6455-v30.xml emd-6455-v30.xml emd-6455.xml emd-6455.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6455_fsc.xml emd_6455_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  400_6455.gif 400_6455.gif 80_6455.gif 80_6455.gif | 59.5 KB 4.2 KB | ||
| Others |  emd_6455_additional_1.map.gz emd_6455_additional_1.map.gz emd_6455_half_map_1.map.gz emd_6455_half_map_1.map.gz emd_6455_half_map_2.map.gz emd_6455_half_map_2.map.gz | 45.9 MB 45.9 MB 45.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6455 http://ftp.pdbj.org/pub/emdb/structures/EMD-6455 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6455 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6455 | HTTPS FTP |
-Validation report
| Summary document |  emd_6455_validation.pdf.gz emd_6455_validation.pdf.gz | 342.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6455_full_validation.pdf.gz emd_6455_full_validation.pdf.gz | 341.7 KB | Display | |
| Data in XML |  emd_6455_validation.xml.gz emd_6455_validation.xml.gz | 10 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6455 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6455 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6455 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6455 | HTTPS FTP |
-Related structure data
| Related structure data |  5an8MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6455.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6455.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TRPV2, sharpened with a B factor of -76 using RELION postprocess | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.31 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
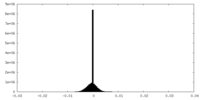
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 6455 additional 1.map
| File | emd_6455_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
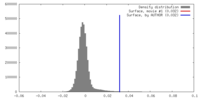
| Density Histograms |
-Supplemental map: emd 6455 half map 1.map
| File | emd_6455_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
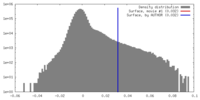
| Density Histograms |
-Supplemental map: emd 6455 half map 2.map
| File | emd_6455_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
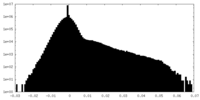
| Density Histograms |
- Sample components
Sample components
-Entire : Rabbit TRPV2
| Entire | Name: Rabbit TRPV2 |
|---|---|
| Components |
|
-Supramolecule #1000: Rabbit TRPV2
| Supramolecule | Name: Rabbit TRPV2 / type: sample / ID: 1000 / Oligomeric state: homotetramer / Number unique components: 1 |
|---|
-Macromolecule #1: TRPV2
| Macromolecule | Name: TRPV2 / type: protein_or_peptide / ID: 1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL |
|---|---|
| Buffer | pH: 7.6 / Details: PBS |
| Grid | Details: 400 mesh C-flat copper grid with carbon support, 1.2 micron holes spaced 1.3 microns |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 120 K / Instrument: GATAN CRYOPLUNGE 3 / Method: Blot for 2.5 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 77 K / Max: 79 K / Average: 78 K |
| Alignment procedure | Legacy - Astigmatism: Objective astigmatism was corrected at 22500 times magnification using Thon rings visualized with a K2 camera. |
| Details | Data was acquired using Leginon and collected in K2 super-resolution mode. |
| Date | May 12, 2015 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 747 / Average electron dose: 57 e/Å2 / Details: 50 frames, 200 ms per frame |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 38168 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 22500 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)