+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5693 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the SecY protein translocation channel in action | |||||||||
 Map data Map data | ribosome-nascent chain-SecYEG complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome-channel complex / active SecYEG channel / nascent chain / E. coli 70S ribosome | |||||||||
| Function / homology |  Function and homology information Function and homology information: / cell envelope Sec protein transport complex / intracellular protein transmembrane transport / protein transport by the Sec complex / protein-transporting ATPase activity / SRP-dependent cotranslational protein targeting to membrane, translocation / signal sequence binding / protein secretion / protein transmembrane transporter activity / negative regulation of translational initiation ...: / cell envelope Sec protein transport complex / intracellular protein transmembrane transport / protein transport by the Sec complex / protein-transporting ATPase activity / SRP-dependent cotranslational protein targeting to membrane, translocation / signal sequence binding / protein secretion / protein transmembrane transporter activity / negative regulation of translational initiation / intracellular protein transport / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding / structural constituent of ribosome / translation / membrane / plasma membrane / cytosol / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 10.1 Å | |||||||||
 Authors Authors | Park E / Menetret JF / Gumbart JC / Ludtke SJ / Li W / Whynot A / Rapoport TA / Akey CW | |||||||||
 Citation Citation |  Journal: Nature / Year: 2014 Journal: Nature / Year: 2014Title: Structure of the SecY channel during initiation of protein translocation. Authors: Eunyong Park / Jean-François Ménétret / James C Gumbart / Steven J Ludtke / Weikai Li / Andrew Whynot / Tom A Rapoport / Christopher W Akey /  Abstract: Many secretory proteins are targeted by signal sequences to a protein-conducting channel, formed by prokaryotic SecY or eukaryotic Sec61 complexes, and are translocated across the membrane during ...Many secretory proteins are targeted by signal sequences to a protein-conducting channel, formed by prokaryotic SecY or eukaryotic Sec61 complexes, and are translocated across the membrane during their synthesis. Crystal structures of the inactive channel show that the SecY subunit of the heterotrimeric complex consists of two halves that form an hourglass-shaped pore with a constriction in the middle of the membrane and a lateral gate that faces the lipid phase. The closed channel has an empty cytoplasmic funnel and an extracellular funnel that is filled with a small helical domain, called the plug. During initiation of translocation, a ribosome-nascent chain complex binds to the SecY (or Sec61) complex, resulting in insertion of the nascent chain. However, the mechanism of channel opening during translocation is unclear. Here we have addressed this question by determining structures of inactive and active ribosome-channel complexes with cryo-electron microscopy. Non-translating ribosome-SecY channel complexes derived from Methanocaldococcus jannaschii or Escherichia coli show the channel in its closed state, and indicate that ribosome binding per se causes only minor changes. The structure of an active E. coli ribosome-channel complex demonstrates that the nascent chain opens the channel, causing mostly rigid body movements of the amino- and carboxy-terminal halves of SecY. In this early translocation intermediate, the polypeptide inserts as a loop into the SecY channel with the hydrophobic signal sequence intercalated into the open lateral gate. The nascent chain also forms a loop on the cytoplasmic surface of SecY rather than entering the channel directly. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5693.map.gz emd_5693.map.gz | 3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5693-v30.xml emd-5693-v30.xml emd-5693.xml emd-5693.xml | 35.5 KB 35.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5693_1.jpg emd_5693_1.jpg | 182.5 KB | ||
| Masks |  emd_5693_msk_1.map emd_5693_msk_1.map emd_5693_msk_2.map emd_5693_msk_2.map emd_5693_msk_3.map emd_5693_msk_3.map emd_5693_msk_4.map emd_5693_msk_4.map emd_5693_msk_5.map emd_5693_msk_5.map emd_5693_msk_6.map emd_5693_msk_6.map emd_5693_msk_7.map emd_5693_msk_7.map emd_5693_msk_8.map emd_5693_msk_8.map emd_5693_msk_9.map emd_5693_msk_9.map | 27 MB 601 KB 27 MB 263.3 KB 27 MB 27 MB 27 MB 27 MB 4.3 MB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5693 http://ftp.pdbj.org/pub/emdb/structures/EMD-5693 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5693 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5693 | HTTPS FTP |
-Validation report
| Summary document |  emd_5693_validation.pdf.gz emd_5693_validation.pdf.gz | 620 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5693_full_validation.pdf.gz emd_5693_full_validation.pdf.gz | 619.5 KB | Display | |
| Data in XML |  emd_5693_validation.xml.gz emd_5693_validation.xml.gz | 5.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5693 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5693 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5693 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5693 | HTTPS FTP |
-Related structure data
| Related structure data |  3j46MC  5691C  5692C  3j45C  4v4nC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5693.map.gz / Format: CCP4 / Size: 26.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5693.map.gz / Format: CCP4 / Size: 26.4 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ribosome-nascent chain-SecYEG complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.12 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
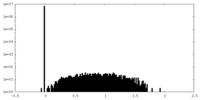
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Segmentation: segmented large ribosomal subunit
| Annotation | segmented large ribosomal subunit | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_1.map emd_5693_msk_1.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
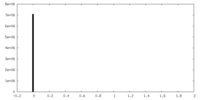
| Density Histograms |
-Segmentation: segmented A and P-site tRNAs
| Annotation | segmented A and P-site tRNAs | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_2.map emd_5693_msk_2.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
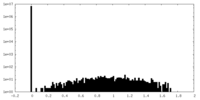
| Density Histograms |
-Segmentation: micelle map from channel density
| Annotation | micelle map from channel density | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_3.map emd_5693_msk_3.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
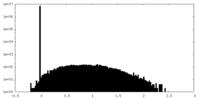
| Density Histograms |
-Segmentation: zoned SecYEG channel
| Annotation | zoned SecYEG channel | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_4.map emd_5693_msk_4.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Segmentation: full channel density with micelle and nascent chain
| Annotation | full channel density with micelle and nascent chain | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_5.map emd_5693_msk_5.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Segmentation: full nascent chain density, excluding fragments in the tunnel
| Annotation | full nascent chain density, excluding fragments in the tunnel | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_6.map emd_5693_msk_6.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Segmentation: nascent chain density with signal sequence helix in the channel
| Annotation | nascent chain density with signal sequence helix in the channel | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_7.map emd_5693_msk_7.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Segmentation: segmented S1 density, excluding flexible fragments
| Annotation | segmented S1 density, excluding flexible fragments | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_8.map emd_5693_msk_8.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Segmentation: segmented small subunit
| Annotation | segmented small subunit | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| File |  emd_5693_msk_9.map emd_5693_msk_9.map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ribosome-nascent chain-SecYEG complex
| Entire | Name: ribosome-nascent chain-SecYEG complex |
|---|---|
| Components |
|
-Supramolecule #1000: ribosome-nascent chain-SecYEG complex
| Supramolecule | Name: ribosome-nascent chain-SecYEG complex / type: sample / ID: 1000 Details: Sample was prepared by crosslinking the nascent chain within stalled membrane associated ribosomes in bacteria, then purified for cryo-EM. Oligomeric state: monomer / Number unique components: 5 |
|---|---|
| Molecular weight | Theoretical: 2.5 MDa |
-Supramolecule #1: membrane-bound 70S ribosome
| Supramolecule | Name: membrane-bound 70S ribosome / type: complex / ID: 1 / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-prokaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 2.5 MDa |
-Macromolecule #1: preprotein translocase
| Macromolecule | Name: preprotein translocase / type: protein_or_peptide / ID: 1 / Name.synonym: SecYEG channel / Details: SecY: P0AGA2, SecE: P0AG96, SecG: P0AG99 / Number of copies: 1 / Oligomeric state: heterotrimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 73.4 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: S1P
| Macromolecule | Name: S1P / type: protein_or_peptide / ID: 2 Name.synonym: endogenous E. coli small ribosomal subunit protein Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 61 KDa |
-Macromolecule #3: transfer RNA
| Macromolecule | Name: transfer RNA / type: rna / ID: 3 / Name.synonym: tRNA Details: Both A- and P-site tRNAs present in purified ribosome-nascent chain-SecYEG complex, stalled with SecM sequence Classification: TRANSFER / Structure: DOUBLE HELIX / Synthetic?: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL |
|---|---|
| Buffer | pH: 7.2 Details: 50 mM Tris-acetate, 10 mM Mg(OAc)2, 80 mM KOAc, 0.06% DDM |
| Grid | Details: 400 mesh Quantifoil holey grids with 2/1 or 1.2/1.2 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 77 K / Instrument: FEI VITROBOT MARK III Details: A homemade freezing device with N2 gas driven plunger was also used to prepare grids in a cabinet to maintain humidity. Method: Blot 1-2 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Temperature | Min: 94 K |
| Alignment procedure | Legacy - Astigmatism: Carbon grain was imaged at ~175000 times magnification and Thon rings were optimized manually, as visualized on a Fourier transform of ccd images. |
| Details | Low dose imaging: automated single particle data collection program from TVIPS was used. |
| Date | Feb 10, 2012 |
| Image recording | Category: CCD / Film or detector model: GENERIC TVIPS (4k x 4k) / Number real images: 4900 / Average electron dose: 20 e/Å2 Details: CCD 4k x 4k image frames: best 4900 from ~5900 images used for data processing. Bits/pixel: 16 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 160 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder: Oxford cold holder / Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | ~450000 particles were selected with e2boxer, used for per ccd frame CTF correction, and then subjected to unsupervised classification to remove aggregates to give 167000 particles. These particles were subjected to supervised classification in 2 steps to give a data set enriched in channels. Particles with the best signal to noise ratio were identified using the FRC comparator from the refinemulti run and classified with e2ligandclassify.py. |
|---|---|
| CTF correction | Details: per micrograph |
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 10.1 Å / Resolution method: OTHER / Software - Name: EMAN2 Details: The structure was solved twice: first with a model starting from a 25-Angstrom filtered E. coli ribosome map generated in house, and then a second time using a filtered ribosome model (EMD- ...Details: The structure was solved twice: first with a model starting from a 25-Angstrom filtered E. coli ribosome map generated in house, and then a second time using a filtered ribosome model (EMD-5036). In each case, after convergence, maps from two EMAN2 refinements with different parameters were averaged after alignment in Chimera. Four maps in total were averaged to reduce the noise. Number images used: 53000 |
| Final two d classification | Number classes: 5300 |
-Atomic model buiding 1
| Initial model | PDB ID:  2i2p |
|---|---|
| Software | Name: Chimera, MDFF |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-3j46: |
-Atomic model buiding 2
| Initial model | PDB ID:  3j01 |
|---|---|
| Software | Name: Chimera, MDFF |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-3j46: |
-Atomic model buiding 3
| Initial model | PDB ID:  3i8g Chain - #0 - Chain ID: B / Chain - #1 - Chain ID: C |
|---|---|
| Software | Name: Chimera, MDFF |
| Details | A- and P-site tRNAs from T. thermophilus ribosome structure. mRNA from this structure also used for some modeling steps. |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-3j46: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)