[English] 日本語
 Yorodumi
Yorodumi- EMDB-3747: Poliovirus type 3 (strain Saukett) stabilized virus-like particle -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3747 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Poliovirus type 3 (strain Saukett) stabilized virus-like particle | |||||||||
 Map data Map data | MRC format map file. Contour level 0.0725 determined by visual inspection of map in UCSF Chimera. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Poliovirus / virus-like particle / vaccine / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationcaveolin-mediated endocytosis of virus by host cell / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane ...caveolin-mediated endocytosis of virus by host cell / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / viral capsid / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / host cell cytoplasm / DNA replication / RNA helicase activity / symbiont-mediated suppression of host gene expression / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / proteolysis / RNA binding / zinc ion binding / ATP binding Similarity search - Function | |||||||||
| Biological species |  Human poliovirus 3 Human poliovirus 3 | |||||||||
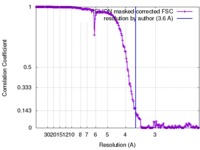
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Bahar MW / Kotecha A | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Plant-made polio type 3 stabilized VLPs-a candidate synthetic polio vaccine. Authors: Johanna Marsian / Helen Fox / Mohammad W Bahar / Abhay Kotecha / Elizabeth E Fry / David I Stuart / Andrew J Macadam / David J Rowlands / George P Lomonossoff /  Abstract: Poliovirus (PV) is the causative agent of poliomyelitis, a crippling human disease known since antiquity. PV occurs in two distinct antigenic forms, D and C, of which only the D form elicits a robust ...Poliovirus (PV) is the causative agent of poliomyelitis, a crippling human disease known since antiquity. PV occurs in two distinct antigenic forms, D and C, of which only the D form elicits a robust neutralizing response. Developing a synthetically produced stabilized virus-like particle (sVLP)-based vaccine with D antigenicity, without the drawbacks of current vaccines, will be a major step towards the final eradication of poliovirus. Such a sVLP would retain the native antigenic conformation and the repetitive structure of the original virus particle, but lack infectious genomic material. In this study, we report the production of synthetically stabilized PV VLPs in plants. Mice carrying the gene for the human PV receptor are protected from wild-type PV when immunized with the plant-made PV sVLPs. Structural analysis of the stabilized mutant at 3.6 Å resolution by cryo-electron microscopy and single-particle reconstruction reveals a structure almost indistinguishable from wild-type PV3.Despite the success of current vaccination against poliomyelitis, safe, cheap and effective vaccines remain sought for continuing eradication effort. Here the authors use plants to express stabilized virus-like particles of type 3 poliovirus that can induce a protective immune response in mice transgenic for the human poliovirus receptor. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3747.map.gz emd_3747.map.gz | 472.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3747-v30.xml emd-3747-v30.xml emd-3747.xml emd-3747.xml | 18.6 KB 18.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3747_fsc.xml emd_3747_fsc.xml | 17.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_3747.png emd_3747.png | 328.5 KB | ||
| Filedesc metadata |  emd-3747.cif.gz emd-3747.cif.gz | 6.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3747 http://ftp.pdbj.org/pub/emdb/structures/EMD-3747 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3747 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3747 | HTTPS FTP |
-Related structure data
| Related structure data |  5o5bMC  3749C  5o5pC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3747.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3747.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | MRC format map file. Contour level 0.0725 determined by visual inspection of map in UCSF Chimera. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Human poliovirus 3
| Entire | Name:  Human poliovirus 3 Human poliovirus 3 |
|---|---|
| Components |
|
-Supramolecule #1: Human poliovirus 3
| Supramolecule | Name: Human poliovirus 3 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all Details: Poliovirus type 3 (Saukett strain) virus-like particle produced in plant expression system. NCBI-ID: 12086 / Sci species name: Human poliovirus 3 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: Yes |
|---|---|
| Host (natural) | Organism:  Human poliovirus 3 / Strain: Saukett Human poliovirus 3 / Strain: Saukett |
| Molecular weight | Theoretical: 5.8 MDa |
| Virus shell | Shell ID: 1 / Name: Capsid / Diameter: 327.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: Capsid proteins, VP1
| Macromolecule | Name: Capsid proteins, VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 3 / Strain: Saukett Human poliovirus 3 / Strain: Saukett |
| Molecular weight | Theoretical: 33.562785 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GIEDLITEVA QGALTLSLPK QQDSLPDTKA SGPAHSKEVP ALTAVETGAT NPLVPSDTVQ TRHVIQRRSR SESTIESFFA RGACVAIIE VDNEEPTTRA QKLFAMWRIT YKDTVQLRRK LEFFTYSRFD MELTFVVTAN FTNTNNGHAL NQVYQIMYIP P GAPTPKSW ...String: GIEDLITEVA QGALTLSLPK QQDSLPDTKA SGPAHSKEVP ALTAVETGAT NPLVPSDTVQ TRHVIQRRSR SESTIESFFA RGACVAIIE VDNEEPTTRA QKLFAMWRIT YKDTVQLRRK LEFFTYSRFD MELTFVVTAN FTNTNNGHAL NQVYQIMYIP P GAPTPKSW DDYTWQTSSN PSIFYTYGAA PARISVPYVG LANAYSHFYD GFAKVPLKTD ANDQIGDSLY SAMTVDDFGV LA IRVVNDH NPTKVTSKVR IYMKPKHVRV WCPRPPRAVP YYGPGVDYKD NLNPLSEKGL TTY UniProtKB: Genome polyprotein |
-Macromolecule #2: Capsid proteins, VP2
| Macromolecule | Name: Capsid proteins, VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 3 / Strain: Saukett Human poliovirus 3 / Strain: Saukett |
| Molecular weight | Theoretical: 30.188982 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SPNVEACGYS DRVLQLTIGN STITTQEAAN SVVAYGRWPE FIRDDEANPV DQPTEPDVAT CRFYTLDTVM WGKESKGWWW KLPDALRDM GLFGQNMYYH YLGRSGYTVH VQCNASKFHQ GALGVFAIPE YCLAGDSDKQ RYTSYANANP GEKGGKFYSQ F NRDTAVTS ...String: SPNVEACGYS DRVLQLTIGN STITTQEAAN SVVAYGRWPE FIRDDEANPV DQPTEPDVAT CRFYTLDTVM WGKESKGWWW KLPDALRDM GLFGQNMYYH YLGRSGYTVH VQCNASKFHQ GALGVFAIPE YCLAGDSDKQ RYTSYANANP GEKGGKFYSQ F NRDTAVTS PKREFCPVDY LLGCGVLLGN AFVYPHQIIN LRTNNSATIV LPYVNAMAID SMVKHNNWGI AILPLSPLDF AQ ESSVEIP ITVTIAPMCS EFNGLRNVTA PKFQ UniProtKB: Genome polyprotein |
-Macromolecule #3: Capsid proteins, VP3
| Macromolecule | Name: Capsid proteins, VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 3 / Strain: Saukett Human poliovirus 3 / Strain: Saukett |
| Molecular weight | Theoretical: 26.3151 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GLPVLNTPGS NQYLTSDNYQ SPCAIPEFDV TPPIDIPGEV KNMMELAEID TMIPLNLENT KRNTMDMYRV TLSDSADLSQ PILCFSLSP ASDPRLSHTM LGEVLNYYTH WAGSLKFTFL FCGSMMATGK ILVAYAPPGA QPPTSRKEAM LGTHVIWDLG L QSSCTMVV ...String: GLPVLNTPGS NQYLTSDNYQ SPCAIPEFDV TPPIDIPGEV KNMMELAEID TMIPLNLENT KRNTMDMYRV TLSDSADLSQ PILCFSLSP ASDPRLSHTM LGEVLNYYTH WAGSLKFTFL FCGSMMATGK ILVAYAPPGA QPPTSRKEAM LGTHVIWDLG L QSSCTMVV PWISNVTYRQ TTQDSFTEGG YISMFYQTRI VVPLSTPKSM SMLGFVSACN DFSVRLLRDT THISQSALPQ UniProtKB: Genome polyprotein |
-Macromolecule #4: Capsid proteins, VP4
| Macromolecule | Name: Capsid proteins, VP4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Human poliovirus 3 / Strain: Saukett Human poliovirus 3 / Strain: Saukett |
| Molecular weight | Theoretical: 7.452113 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGAQVSSQKV GAHENSNRAY GGSTINYTTI NYYKDSASNA ASKQDYSQDP SKFTEPLKDV LIKTAPALN UniProtKB: Genome polyprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL |
|---|---|
| Buffer | pH: 7 / Details: PBS pH 7.0 |
| Grid | Model: C-flat CF-2/1-2C / Material: COPPER / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK I / Details: Blot for 4 seconds before plunging. |
| Details | Virus-like particles for polio type 3 (strain Saukett) were purified by Nycodenz gradients and assessed for monodispersity by negative stain EM analysis. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4086 pixel / Digitization - Dimensions - Height: 4086 pixel / Digitization - Frames/image: 2-33 / Number grids imaged: 1 / Number real images: 2768 / Average exposure time: 0.2 sec. / Average electron dose: 1.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 133333 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | phenix.real_space_refine was used to refine the atomic model in the cryo-em map. |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: Cross-correlation coefficient |
| Output model |  PDB-5o5b: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)