+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of ATG2A-WIPI4 complex | |||||||||
 Map data Map data | Half map A of ATG2A-WIPI4 complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ATG2A / single particle cryo-EM / peripheral membrane proteins / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationphagophore / lipid transfer activity / organelle membrane contact site / glycophagy / positive regulation of autophagosome assembly / phagophore assembly site membrane / piecemeal microautophagy of the nucleus / phosphatidylinositol-3-phosphate binding / phagophore assembly site / reticulophagy ...phagophore / lipid transfer activity / organelle membrane contact site / glycophagy / positive regulation of autophagosome assembly / phagophore assembly site membrane / piecemeal microautophagy of the nucleus / phosphatidylinositol-3-phosphate binding / phagophore assembly site / reticulophagy / autophagosome assembly / protein-membrane adaptor activity / lipid droplet / endoplasmic reticulum membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
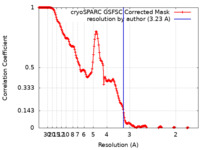
| Method | single particle reconstruction / cryo EM / Resolution: 3.23 Å | |||||||||
 Authors Authors | Wang Y / Stjepanovic G | |||||||||
| Funding support |  China, 2 items China, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structural basis for lipid transfer by the ATG2A-ATG9A complex Authors: Wang Y / Stjepanovic G | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_37087.map.gz emd_37087.map.gz | 483.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-37087-v30.xml emd-37087-v30.xml emd-37087.xml emd-37087.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_37087_fsc.xml emd_37087_fsc.xml | 17 KB | Display |  FSC data file FSC data file |
| Images |  emd_37087.png emd_37087.png | 55.5 KB | ||
| Filedesc metadata |  emd-37087.cif.gz emd-37087.cif.gz | 6.3 KB | ||
| Others |  emd_37087_half_map_1.map.gz emd_37087_half_map_1.map.gz emd_37087_half_map_2.map.gz emd_37087_half_map_2.map.gz | 475.8 MB 475.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-37087 http://ftp.pdbj.org/pub/emdb/structures/EMD-37087 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37087 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-37087 | HTTPS FTP |
-Validation report
| Summary document |  emd_37087_validation.pdf.gz emd_37087_validation.pdf.gz | 757.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_37087_full_validation.pdf.gz emd_37087_full_validation.pdf.gz | 757.2 KB | Display | |
| Data in XML |  emd_37087_validation.xml.gz emd_37087_validation.xml.gz | 26.6 KB | Display | |
| Data in CIF |  emd_37087_validation.cif.gz emd_37087_validation.cif.gz | 34.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37087 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37087 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37087 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-37087 | HTTPS FTP |
-Related structure data
| Related structure data |  8kbyMC  8kbxC  8kbzC  8kc3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_37087.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_37087.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A of ATG2A-WIPI4 complex | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: Half map B of ATG2A-WIPI4 complex
| File | emd_37087_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B of ATG2A-WIPI4 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B of ATG2A-WIPI4 complex
| File | emd_37087_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B of ATG2A-WIPI4 complex | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : ATG2A
| Entire | Name: ATG2A |
|---|---|
| Components |
|
-Supramolecule #1: ATG2A
| Supramolecule | Name: ATG2A / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 213 KDa |
-Macromolecule #1: Autophagy-related protein 2 homolog A
| Macromolecule | Name: Autophagy-related protein 2 homolog A / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 213.100281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSRWLWPWSN CVKERVCRYL LHHYLGHFFQ EHLSLDQLSL DLYKGSVALR DIHLEIWSVN EVLESMESPL ELVEGFVGSI EVAVPWAAL LTDHCTVRVS GLQLTLQPRR GPAPGAADSQ SWASCMTTSL QLAQECLRDG LPEPSEPPQP LEGLEMFAQT I ETVLRRIK ...String: MSRWLWPWSN CVKERVCRYL LHHYLGHFFQ EHLSLDQLSL DLYKGSVALR DIHLEIWSVN EVLESMESPL ELVEGFVGSI EVAVPWAAL LTDHCTVRVS GLQLTLQPRR GPAPGAADSQ SWASCMTTSL QLAQECLRDG LPEPSEPPQP LEGLEMFAQT I ETVLRRIK VTFLDTVVRV EHSPGDGERG VAVEVRVQRL EYCDEAVRDP SQAPPVDVHQ PPAFLHKLLQ LAGVRLHYEE LP AQEEPPE PPLQIGSCSG YMELMVKLKQ NEAFPGPKLE VAGQLGSLHL LLTPRQLQQL QELLSAVSLT DHEGLADKLN KSR PLGAED LWLIEQDLNQ QLQAGAVAEP LSPDPLTNPL LNLDNTDLFF SMAGLTSSVA SALSELSLSD VDLASSVRSD MASR RLSAQ AHPAGKMAPN PLLDTMRPDS LLKMTLGGVT LTLLQTSAPS SGPPDLATHF FTEFDATKDG PFGSRDFHHL RPRFQ RACP CSHVRLTGTA VQLSWELRTG SRGRRTTSME VHFGQLEVLE CLWPRGTSEP EYTEILTFPG TLGSQASARP CAHLRH TQI LRRVPKSRPR RSVACHCHSE LALDLANFQA DVELGALDRL AALLRLATVP AEPPAGLLTE PLPAMEQQTV FRLSAPR AT LRLRFPIADL RPEPDPWAGQ AVRAEQLRLE LSEPQFRSEL SSGPGPPVPT HLELTCSDLH GIYEDGGKPP VPCLRVSK A LDPKSTGRKY FLPQVVVTVN PQSSSTQWEV APEKGEELEL SVESPCELRE PEPSPFSSKR TMYETEEMVI PGDPEEMRT FQSRTLALSR CSLEVILPSV HIFLPSKEVY ESIYNRINND LLMWEPADLL PTPDPAAQPS GFPGPSGFWH DSFKMCKSAF KLANCFDLT PDSDSDDEDA HFFSVGASGG PQAAAPEAPS LHLQSTFSTL VTVLKGRITA LCETKDEGGK RLEAVHGELV L DMEHGTLF SVSQYCGQPG LGYFCLEAEK ATLYHRAAVD DYPLPSHLDL PSFAPPAQLA PTIYPSEEGV TERGASGRKG QG RGPHMLS TAVRIHLDPH KNVKEFLVTL RLHKATLRHY MALPEQSWHS QLLEFLDVLD DPVLGYLPPT VITILHTHLF SCS VDYRPL YLPVRVLITA ETFTLSSNII MDTSTFLLRF ILDDSALYLS DKCEVETLDL RRDYVCVLDV DLLELVIKTW KGST EGKLS QPLFELRCSN NVVHVHSCAD SCALLVNLLQ YVMSTGDLHP PPRPPSPTEI AGQKLSESPA SLPSCPPVET ALINQ RDLA DALLDTERSL RELAQPSGGH LPQASPISVY LFPGERSGAP PPSPPVGGPA GSLGSCSEEK EDEREEEGDG DTLDSD EFC ILDAPGLGIP PRDGEPVVTQ LHPGPIVVRD GYFSRPIGST DLLRAPAHFP VPSTRVVLRE VSLVWHLYGG RDFGPHP GH RARTGLSGPR SSPSRCSGPN RPQNSWRTQG GSGRQHHVLM EIQLSKVSFQ HEVYPAEPAT GPAAPSQELE ERPLSRQV F IVQELEVRDR LASSQINKFL YLHTSERMPR RAHSNMLTIK ALHVAPTTNL GGPECCLRVS LMPLRLNVDQ DALFFLKDF FTSLVAGINP VVPGETSAEA RPETRAQPSS PLEGQAEGVE TTGSQEAPGG GHSPSPPDQQ PIYFREFRFT SEVPIWLDYH GKHVTMDQV GTFAGLLIGL AQLNCSELKL KRLCCRHGLL GVDKVLGYAL NEWLQDIRKN QLPGLLGGVG PMHSVVQLFQ G FRDLLWLP IEQYRKDGRL MRGLQRGAAS FGSSTASAAL ELSNRLVQAI QATAETVYDI LSPAAPVSRS LQDKRSARRL RR GQQPADL REGVAKAYDT VREGILDTAQ TICDVASRGH EQKGLTGAVG GVIRQLPPTV VKPLILATEA TSSLLGGMRN QIV PDAHKD HALKWRSDSA QD UniProtKB: Autophagy-related protein 2 homolog A |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 61.65 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 105000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z
Z Y
Y X
X