[English] 日本語
 Yorodumi
Yorodumi- EMDB-30979: SARS-CoV-2 spike in complex with the Ab1 neutralizing antibody (f... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30979 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 spike in complex with the Ab1 neutralizing antibody (focused refinement on Fab-RBD) | |||||||||
 Map data Map data | focused refinement of RBD-Fab sub-complex (S-Ab1) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Spike / SARS-CoV-2 / Antibody / VIRAL PROTEIN / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / viral translation / host extracellular space / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / viral translation / host extracellular space / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / entry receptor-mediated virion attachment to host cell / membrane fusion / Attachment and Entry / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / receptor ligand activity / endocytosis involved in viral entry into host cell / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
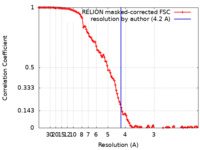
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Liu C | |||||||||
 Citation Citation |  Journal: Cell Discov / Year: 2021 Journal: Cell Discov / Year: 2021Title: Three epitope-distinct human antibodies from RenMab mice neutralize SARS-CoV-2 and cooperatively minimize the escape of mutants. Authors: Jianhui Nie / Jingshu Xie / Shuo Liu / Jiajing Wu / Chuan Liu / Jianhui Li / Yacui Liu / Meiyu Wang / Huizhen Zhao / Yabo Zhang / Jiawei Yao / Lei Chen / Yuelei Shen / Yi Yang / Hong-Wei ...Authors: Jianhui Nie / Jingshu Xie / Shuo Liu / Jiajing Wu / Chuan Liu / Jianhui Li / Yacui Liu / Meiyu Wang / Huizhen Zhao / Yabo Zhang / Jiawei Yao / Lei Chen / Yuelei Shen / Yi Yang / Hong-Wei Wang / Youchun Wang / Weijin Huang /  Abstract: Coronavirus disease 2019 (COVID-19), a pandemic disease caused by the newly emerging severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has caused more than 3.8 million deaths to date. ...Coronavirus disease 2019 (COVID-19), a pandemic disease caused by the newly emerging severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has caused more than 3.8 million deaths to date. Neutralizing antibodies are effective therapeutic measures. However, many naturally occurring mutations at the receptor-binding domain (RBD) have emerged, and some of them can evade existing neutralizing antibodies. Here, we utilized RenMab, a novel mouse carrying the entire human antibody variable region, for neutralizing antibody discovery. We obtained several potent RBD-blocking antibodies and categorized them into four distinct groups by epitope mapping. We determined the involved residues of the epitope of three representative antibodies by cryo-electron microscopy (Cryo-EM) studies. Moreover, we performed neutralizing experiments with 50 variant strains with single or combined mutations and found that the mixing of three epitope-distinct antibodies almost eliminated the mutant escape. Our study provides a sound basis for the rational design of fully human antibody cocktails against SARS-CoV-2 and pre-emergent coronaviral threats. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30979.map.gz emd_30979.map.gz | 5.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30979-v30.xml emd-30979-v30.xml emd-30979.xml emd-30979.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_30979_fsc.xml emd_30979_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_30979.png emd_30979.png | 77.9 KB | ||
| Filedesc metadata |  emd-30979.cif.gz emd-30979.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30979 http://ftp.pdbj.org/pub/emdb/structures/EMD-30979 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30979 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30979 | HTTPS FTP |
-Validation report
| Summary document |  emd_30979_validation.pdf.gz emd_30979_validation.pdf.gz | 344.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_30979_full_validation.pdf.gz emd_30979_full_validation.pdf.gz | 344.1 KB | Display | |
| Data in XML |  emd_30979_validation.xml.gz emd_30979_validation.xml.gz | 11.9 KB | Display | |
| Data in CIF |  emd_30979_validation.cif.gz emd_30979_validation.cif.gz | 16.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30979 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30979 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30979 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-30979 | HTTPS FTP |
-Related structure data
| Related structure data |  7e3cMC  7e39C  7e3bC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30979.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30979.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | focused refinement of RBD-Fab sub-complex (S-Ab1) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.087 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : complex of spike protein trimer with three Ab1 antibodies
| Entire | Name: complex of spike protein trimer with three Ab1 antibodies |
|---|---|
| Components |
|
-Supramolecule #1: complex of spike protein trimer with three Ab1 antibodies
| Supramolecule | Name: complex of spike protein trimer with three Ab1 antibodies type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|
-Supramolecule #2: Spike protein S1
| Supramolecule | Name: Spike protein S1 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: Light chain of Ab1
| Supramolecule | Name: Light chain of Ab1 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #4: Heavy chain of Ab1
| Supramolecule | Name: Heavy chain of Ab1 / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike protein S1
| Macromolecule | Name: Spike protein S1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 21.776381 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT ...String: TNLCPFGEVF NATRFASVYA WNRKRISNCV ADYSVLYNSA SFSTFKCYGV SPTKLNDLCF TNVYADSFVI RGDEVRQIAP GQTGKIADY NYKLPDDFTG CVIAWNSNNL DSKVGGNYNY LYRLFRKSNL KPFERDISTE IYQAGSTPCN GVEGFNCYFP L QSYGFQPT NGVGYQPYRV VVLSFELLHA PATVCG UniProtKB: Spike glycoprotein |
-Macromolecule #2: Light chain of Ab1
| Macromolecule | Name: Light chain of Ab1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.417027 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQLTQSPSF LSASVGDRVT ITCRASQGIS NYLAWYQQKP GKAPKLLIYA ASTLQTGVPS RFSGSGSGTE FTLTISSLQP EDFATYYCQ QLNSYPPLTF GGGSRVEIKR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: DIQLTQSPSF LSASVGDRVT ITCRASQGIS NYLAWYQQKP GKAPKLLIYA ASTLQTGVPS RFSGSGSGTE FTLTISSLQP EDFATYYCQ QLNSYPPLTF GGGSRVEIKR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGEC |
-Macromolecule #3: Heavy chain of Ab1
| Macromolecule | Name: Heavy chain of Ab1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 48.707785 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVLLVESGGG LVQPGGSLRL SCAASGLTVS SNYMTWVRQA PGKGLEWVSV IYSGGSTFYA DSVKGRCTIS RHNSKNTLYL QMNSLRAED TAVYYCARDL DYYGMDVWGQ GTTVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY FPEPVTVSWN S GALTSGVH ...String: EVLLVESGGG LVQPGGSLRL SCAASGLTVS SNYMTWVRQA PGKGLEWVSV IYSGGSTFYA DSVKGRCTIS RHNSKNTLYL QMNSLRAED TAVYYCARDL DYYGMDVWGQ GTTVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY FPEPVTVSWN S GALTSGVH TFPAVLQSSG LYSLSSVVTV PSSSLGTQTY ICNVNHKPSN TKVDKKVEPK SCDKTHTCPP CPAPEAAGGP SV FLFPPKP KDTLMISRTP EVTCVVVDVS HEDPEVKFNW YVDGVEVHNA KTKPREEQYN STYRVVSVLT VLHQDWLNGK EYK CKVSNK ALPAPIEKTI SKAKGQPREP QVYTLPPSRE EMTKNQVSLT CLVKGFYPSD IAVEWESNGQ PENNYKTTPP VLDS DGSFF LYSKLTVDKS RWQQGNVFSC SVMHEALHNH YTQKSLSLSP GK |
-Macromolecule #4: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 4 / Number of copies: 1 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)