+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | P. polymyxa GS(12) - apo | ||||||||||||
 Map data Map data | Sharpened (B factor 92 A^2) | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | glutamine synthetase dodecamer / BIOSYNTHETIC PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationpolyamine catabolic process / glutamine synthetase / glutamine biosynthetic process / glutamine synthetase activity / ATP binding / metal ion binding / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  Paenibacillus polymyxa (bacteria) Paenibacillus polymyxa (bacteria) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.16 Å | ||||||||||||
 Authors Authors | Travis BA / Peck J | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Molecular dissection of the glutamine synthetase-GlnR nitrogen regulatory circuitry in Gram-positive bacteria. Authors: Brady A Travis / Jared V Peck / Raul Salinas / Brandon Dopkins / Nicholas Lent / Viet D Nguyen / Mario J Borgnia / Richard G Brennan / Maria A Schumacher /  Abstract: How bacteria sense and respond to nitrogen levels are central questions in microbial physiology. In Gram-positive bacteria, nitrogen homeostasis is controlled by an operon encoding glutamine ...How bacteria sense and respond to nitrogen levels are central questions in microbial physiology. In Gram-positive bacteria, nitrogen homeostasis is controlled by an operon encoding glutamine synthetase (GS), a dodecameric machine that assimilates ammonium into glutamine, and the GlnR repressor. GlnR detects nitrogen excess indirectly by binding glutamine-feedback-inhibited-GS (FBI-GS), which activates its transcription-repression function. The molecular mechanisms behind this regulatory circuitry, however, are unknown. Here we describe biochemical and structural analyses of GS and FBI-GS-GlnR complexes from pathogenic and non-pathogenic Gram-positive bacteria. The structures show FBI-GS binds the GlnR C-terminal domain within its active-site cavity, juxtaposing two GlnR monomers to form a DNA-binding-competent GlnR dimer. The FBI-GS-GlnR interaction stabilizes the inactive GS conformation. Strikingly, this interaction also favors a remarkable dodecamer to tetradecamer transition in some GS, breaking the paradigm that all bacterial GS are dodecamers. These data thus unveil unique structural mechanisms of transcription and enzymatic regulation. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25870.map.gz emd_25870.map.gz | 230.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25870-v30.xml emd-25870-v30.xml emd-25870.xml emd-25870.xml | 13.6 KB 13.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_25870.png emd_25870.png | 113.9 KB | ||
| Filedesc metadata |  emd-25870.cif.gz emd-25870.cif.gz | 5.5 KB | ||
| Others |  emd_25870_additional_1.map.gz emd_25870_additional_1.map.gz | 121.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25870 http://ftp.pdbj.org/pub/emdb/structures/EMD-25870 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25870 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25870 | HTTPS FTP |
-Related structure data
| Related structure data |  7tfdMC  7tdpC  7tdvC  7teaC  7tecC  7tenC  7tf6C  7tf7C  7tf9C  7tfaC  7tfbC  7tfcC  7tfeC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_25870.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25870.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened (B factor 92 A^2) | ||||||||||||||||||||
| Voxel size | X=Y=Z: 0.88 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unsharpened
| File | emd_25870_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
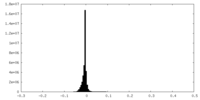
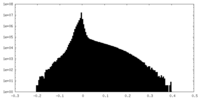
| Density Histograms |
- Sample components
Sample components
-Entire : Apo dodecameric P. polymyxa GS complex
| Entire | Name: Apo dodecameric P. polymyxa GS complex |
|---|---|
| Components |
|
-Supramolecule #1: Apo dodecameric P. polymyxa GS complex
| Supramolecule | Name: Apo dodecameric P. polymyxa GS complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Paenibacillus polymyxa (bacteria) Paenibacillus polymyxa (bacteria) |
-Macromolecule #1: Glutamine synthetase
| Macromolecule | Name: Glutamine synthetase / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: glutamine synthetase |
|---|---|
| Source (natural) | Organism:  Paenibacillus polymyxa (bacteria) Paenibacillus polymyxa (bacteria) |
| Molecular weight | Theoretical: 52.51034 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH MSYTREDIIR IAEEENVRFI RLQFTDLLGT IKNVEIPVSQ LEKALDNKMM FDGSSIEGYV RIEESDMYL YPDLDTWVVF PWVTSDRVAR LICDIYKPDG SPFAGDPRGI LKRVLKEAEE LGYTSMNVGP EPEFFLFKTD E KGDPTTEL ...String: MGSSHHHHHH SSGLVPRGSH MSYTREDIIR IAEEENVRFI RLQFTDLLGT IKNVEIPVSQ LEKALDNKMM FDGSSIEGYV RIEESDMYL YPDLDTWVVF PWVTSDRVAR LICDIYKPDG SPFAGDPRGI LKRVLKEAEE LGYTSMNVGP EPEFFLFKTD E KGDPTTEL NDQGGYFDLA PMDLGENCRR EIVLKLEEMG FEIEASHHEV APGQHEIDFK YADAVKAADQ IQTFKLVVKT IA RQHGLHA TFMPKPLFGV NGSGMHCNQS LFKDNENVFY DETDELGLSQ TARHYMAGIL KHARAMAAIT NPTVNSYKRL VPG YEAPCY VAWSASNRSP MIRIPASRGL STRVEVRNPD PAANPYLALA VMLRAGLDGI KRQMALPAPI DRNIYVMSEE ERIE EGIPS LPADLKEALS ELIRSEVISD ALGDHALAYF YELKEIEWDM YRTQVHQWER DQYLTLY UniProtKB: Glutamine synthetase |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 24 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.9 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 41.67 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: D6 (2x6 fold dihedral) / Resolution.type: BY AUTHOR / Resolution: 3.16 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 3.2) / Number images used: 105664 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3.2) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3.2) |
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-7tfd: |
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X