+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24518 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human p97-R155H mutant bound to ADP. | |||||||||
 Map data Map data | Cryo-EM structure of human full-length p97-R155H mutant bound to ADP. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | p97 / VCP / TERA / Inhibitor / CB-5083 / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationflavin adenine dinucleotide catabolic process / VCP-NSFL1C complex / endoplasmic reticulum stress-induced pre-emptive quality control / endosome to lysosome transport via multivesicular body sorting pathway / BAT3 complex binding / cytoplasmic ubiquitin ligase complex / cellular response to arsenite ion / protein-DNA covalent cross-linking repair / Derlin-1 retrotranslocation complex / positive regulation of protein K63-linked deubiquitination ...flavin adenine dinucleotide catabolic process / VCP-NSFL1C complex / endoplasmic reticulum stress-induced pre-emptive quality control / endosome to lysosome transport via multivesicular body sorting pathway / BAT3 complex binding / cytoplasmic ubiquitin ligase complex / cellular response to arsenite ion / protein-DNA covalent cross-linking repair / Derlin-1 retrotranslocation complex / positive regulation of protein K63-linked deubiquitination / deubiquitinase activator activity / cytoplasm protein quality control / positive regulation of oxidative phosphorylation / aggresome assembly / ubiquitin-modified protein reader activity / regulation of protein localization to chromatin / mitotic spindle disassembly / VCP-NPL4-UFD1 AAA ATPase complex / cellular response to misfolded protein / positive regulation of mitochondrial membrane potential / vesicle-fusing ATPase / K48-linked polyubiquitin modification-dependent protein binding / NAD+ metabolic process / regulation of aerobic respiration / retrograde protein transport, ER to cytosol / stress granule disassembly / ATPase complex / ubiquitin-specific protease binding / regulation of synapse organization / ciliary transition zone / positive regulation of ATP biosynthetic process / intracellular membrane-bounded organelle / ubiquitin-like protein ligase binding / RHOH GTPase cycle / MHC class I protein binding / autophagosome maturation / HSF1 activation / negative regulation of hippo signaling / endoplasmic reticulum to Golgi vesicle-mediated transport / polyubiquitin modification-dependent protein binding / interstrand cross-link repair / ATP metabolic process / translesion synthesis / Attachment and Entry / Protein methylation / negative regulation of protein localization to chromatin / endoplasmic reticulum unfolded protein response / ERAD pathway / proteasomal protein catabolic process / lipid droplet / ciliary tip / proteasome complex / viral genome replication / Josephin domain DUBs / macroautophagy / negative regulation of smoothened signaling pathway / N-glycan trimming in the ER and Calnexin/Calreticulin cycle / establishment of protein localization / positive regulation of protein-containing complex assembly / Hh mutants are degraded by ERAD / Hedgehog ligand biogenesis / Translesion Synthesis by POLH / Defective CFTR causes cystic fibrosis / ADP binding / positive regulation of non-canonical NF-kappaB signal transduction / ABC-family protein mediated transport / autophagy / cytoplasmic stress granule / Aggrephagy / positive regulation of protein catabolic process / azurophil granule lumen / Ovarian tumor domain proteases / KEAP1-NFE2L2 pathway / positive regulation of canonical Wnt signaling pathway / double-strand break repair / positive regulation of proteasomal ubiquitin-dependent protein catabolic process / cellular response to heat / E3 ubiquitin ligases ubiquitinate target proteins / site of double-strand break / Neddylation / secretory granule lumen / protein phosphatase binding / regulation of apoptotic process / ficolin-1-rich granule lumen / ubiquitin-dependent protein catabolic process / Attachment and Entry / proteasome-mediated ubiquitin-dependent protein catabolic process / ciliary basal body / protein ubiquitination / protein domain specific binding / DNA repair / DNA damage response / Neutrophil degranulation / ubiquitin protein ligase binding / lipid binding / endoplasmic reticulum membrane / perinuclear region of cytoplasm / glutamatergic synapse / endoplasmic reticulum / ATP hydrolysis activity Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Caffrey B / Zhu X / Berezuk A / Tuttle K / Chittori S / Subramaniam S | |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2021 Journal: J Biol Chem / Year: 2021Title: AAA+ ATPase p97/VCP mutants and inhibitor binding disrupt inter-domain coupling and subsequent allosteric activation. Authors: Brian Caffrey / Xing Zhu / Alison Berezuk / Katharine Tuttle / Sagar Chittori / Sriram Subramaniam /  Abstract: The human AAA+ ATPase p97, also known as valosin-containing protein, a potential target for cancer therapeutics, plays a vital role in the clearing of misfolded proteins. p97 dysfunction is also ...The human AAA+ ATPase p97, also known as valosin-containing protein, a potential target for cancer therapeutics, plays a vital role in the clearing of misfolded proteins. p97 dysfunction is also known to play a crucial role in several neurodegenerative disorders, such as MultiSystem Proteinopathy 1 (MSP-1) and Familial Amyotrophic Lateral Sclerosis (ALS). However, the structural basis of its role in such diseases remains elusive. Here, we present cryo-EM structural analyses of four disease mutants p97, p97, p97, p97, as well as p97, implicated in resistance to the drug CB-5083, a potent p97 inhibitor. Our cryo-EM structures demonstrate that these mutations affect nucleotide-driven allosteric activation across the three principal p97 domains (N, D1, and D2) by predominantly interfering with either (1) the coupling between the D1 and N-terminal domains (p97 and p97), (2) the interprotomer interactions (p97), or (3) the coupling between D1 and D2 nucleotide domains (p97, p97). We also show that binding of the competitive inhibitor, CB-5083, to the D2 domain prevents conformational changes similar to those seen for mutations that affect coupling between the D1 and D2 domains. Our studies enable tracing of the path of allosteric activation across p97 and establish a common mechanistic link between active site inhibition and defects in allosteric activation by disease-causing mutations and have potential implications for the design of novel allosteric compounds that can modulate p97 function. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24518.map.gz emd_24518.map.gz | 31.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24518-v30.xml emd-24518-v30.xml emd-24518.xml emd-24518.xml | 12.1 KB 12.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24518_fsc.xml emd_24518_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_24518.png emd_24518.png | 60.3 KB | ||
| Filedesc metadata |  emd-24518.cif.gz emd-24518.cif.gz | 5.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24518 http://ftp.pdbj.org/pub/emdb/structures/EMD-24518 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24518 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24518 | HTTPS FTP |
-Related structure data
| Related structure data |  7rl6MC  7rl7C  7rl9C  7rlaC  7rlbC  7rlcC  7rldC  7rlfC  7rlgC  7rlhC  7rliC  7rljC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24518.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24518.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of human full-length p97-R155H mutant bound to ADP. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0125 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
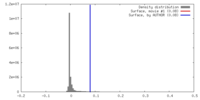
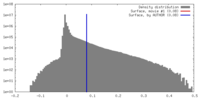
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Full-length Hexameric p97-R155H mutant
| Entire | Name: Full-length Hexameric p97-R155H mutant |
|---|---|
| Components |
|
-Supramolecule #1: Full-length Hexameric p97-R155H mutant
| Supramolecule | Name: Full-length Hexameric p97-R155H mutant / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 540 KDa |
-Macromolecule #1: Transitional endoplasmic reticulum ATPase
| Macromolecule | Name: Transitional endoplasmic reticulum ATPase / type: protein_or_peptide / ID: 1 / Number of copies: 6 / Enantiomer: LEVO / EC number: vesicle-fusing ATPase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 91.472969 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKHHHHHHGT SENLYFQGAS GADSKGDDLS TAILKQKNRP NRLIVDEAIN EDNSVVSLSQ PKMDELQLFR GDTVLLKGKK RREAVCIVL SDDTCSDEKI RMNRVVRNNL RVRLGDVISI QPCPDVKYGK RIHVLPIDDT VEGITGNLFE VYLKPYFLEA Y RPIRKGDI ...String: MKHHHHHHGT SENLYFQGAS GADSKGDDLS TAILKQKNRP NRLIVDEAIN EDNSVVSLSQ PKMDELQLFR GDTVLLKGKK RREAVCIVL SDDTCSDEKI RMNRVVRNNL RVRLGDVISI QPCPDVKYGK RIHVLPIDDT VEGITGNLFE VYLKPYFLEA Y RPIRKGDI FLVHGGMRAV EFKVVETDPS PYCIVAPDTV IHCEGEPIKR EDEEESLNEV GYDDIGGCRK QLAQIKEMVE LP LRHPALF KAIGVKPPRG ILLYGPPGTG KTLIARAVAN ETGAFFFLIN GPEIMSKLAG ESESNLRKAF EEAEKNAPAI IFI DELDAI APKREKTHGE VERRIVSQLL TLMDGLKQRA HVIVMAATNR PNSIDPALRR FGRFDREVDI GIPDATGRLE ILQI HTKNM KLADDVDLEQ VANETHGHVG ADLAALCSEA ALQAIRKKMD LIDLEDETID AEVMNSLAVT MDDFRWALSQ SNPSA LRET VVEVPQVTWE DIGGLEDVKR ELQELVQYPV EHPDKFLKFG MTPSKGVLFY GPPGCGKTLL AKAIANECQA NFISIK GPE LLTMWFGESE ANVREIFDKA RQAAPCVLFF DELDSIAKAR GGNIGDGGGA ADRVINQILT EMDGMSTKKN VFIIGAT NR PDIIDPAILR PGRLDQLIYI PLPDEKSRVA ILKANLRKSP VAKDVDLEFL AKMTNGFSGA DLTEICQRAC KLAIRESI E SEIRRERERQ TNPSAMEVEE DDPVPEIRRD HFEEAMRFAR RSVSDNDIRK YEMFAQTLQQ SRGFGSFRFP SGNQGGAGP SQGSGGGTGG SVYTEDNDDD LYG UniProtKB: Transitional endoplasmic reticulum ATPase |
-Macromolecule #2: ADENOSINE-5'-DIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-DIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 12 / Formula: ADP |
|---|---|
| Molecular weight | Theoretical: 427.201 Da |
| Chemical component information |  ChemComp-ADP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | 2D array |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: Protein Storage Buffer with ADP. | |||||||||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 4410 / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)