[English] 日本語
 Yorodumi
Yorodumi- EMDB-24456: Mycobacterial CIII2CIV2 supercomplex, inhibitor free, -Lpqe cyt c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24456 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mycobacterial CIII2CIV2 supercomplex, inhibitor free, -Lpqe cyt cc open | ||||||||||||
 Map data Map data | Mycobacterium CIII2CIV2 supercomplex, inhibitor free, -Lpqe cyt. cc open. | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationaerobic electron transport chain / cytochrome-c oxidase / quinol-cytochrome-c reductase / ubiquinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / superoxide dismutase / superoxide dismutase activity / : / respiratory electron transport chain / electron transport chain ...aerobic electron transport chain / cytochrome-c oxidase / quinol-cytochrome-c reductase / ubiquinol-cytochrome-c reductase activity / cytochrome-c oxidase activity / superoxide dismutase / superoxide dismutase activity / : / respiratory electron transport chain / electron transport chain / 2 iron, 2 sulfur cluster binding / iron ion binding / copper ion binding / heme binding / membrane / metal ion binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Mycolicibacterium smegmatis MC2 155 (bacteria) / Mycolicibacterium smegmatis MC2 155 (bacteria) /  Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||||||||
 Authors Authors | Di Trani JM / Yanofsky DJ / Rubinstein JL | ||||||||||||
| Funding support |  Canada, Canada,  Sweden, 3 items Sweden, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2021 Journal: Elife / Year: 2021Title: Structure of mycobacterial CIIICIV respiratory supercomplex bound to the tuberculosis drug candidate telacebec (Q203). Authors: David J Yanofsky / Justin M Di Trani / Sylwia Król / Rana Abdelaziz / Stephanie A Bueler / Peter Imming / Peter Brzezinski / John L Rubinstein /    Abstract: The imidazopyridine telacebec, also known as Q203, is one of only a few new classes of compounds in more than 50 years with demonstrated antituberculosis activity in humans. Telacebec inhibits the ...The imidazopyridine telacebec, also known as Q203, is one of only a few new classes of compounds in more than 50 years with demonstrated antituberculosis activity in humans. Telacebec inhibits the mycobacterial respiratory supercomplex composed of complexes III and IV (CIIICIV). In mycobacterial electron transport chains, CIIICIV replaces canonical CIII and CIV, transferring electrons from the intermediate carrier menaquinol to the final acceptor, molecular oxygen, while simultaneously transferring protons across the inner membrane to power ATP synthesis. We show that telacebec inhibits the menaquinol:oxygen oxidoreductase activity of purified CIIICIV at concentrations similar to those needed to inhibit electron transfer in mycobacterial membranes and growth in culture. We then used electron cryomicroscopy (cryoEM) to determine structures of CIIICIV both in the presence and absence of telacebec. The structures suggest that telacebec prevents menaquinol oxidation by blocking two different menaquinol binding modes to prevent CIIICIV activity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24456.map.gz emd_24456.map.gz | 10 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24456-v30.xml emd-24456-v30.xml emd-24456.xml emd-24456.xml | 25.9 KB 25.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24456.png emd_24456.png | 73.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24456 http://ftp.pdbj.org/pub/emdb/structures/EMD-24456 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24456 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24456 | HTTPS FTP |
-Validation report
| Summary document |  emd_24456_validation.pdf.gz emd_24456_validation.pdf.gz | 351.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_24456_full_validation.pdf.gz emd_24456_full_validation.pdf.gz | 350.8 KB | Display | |
| Data in XML |  emd_24456_validation.xml.gz emd_24456_validation.xml.gz | 6.8 KB | Display | |
| Data in CIF |  emd_24456_validation.cif.gz emd_24456_validation.cif.gz | 7.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24456 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24456 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24456 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24456 | HTTPS FTP |
-Related structure data
| Related structure data |  7rh6MC  7rh5C  7rh7C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24456.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24456.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mycobacterium CIII2CIV2 supercomplex, inhibitor free, -Lpqe cyt. cc open. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.03 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
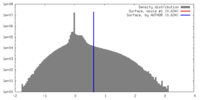
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Respiratory Complex CIII2CIV2SOD2 from Mycobacterium smegmatis
+Supramolecule #1: Respiratory Complex CIII2CIV2SOD2 from Mycobacterium smegmatis
+Macromolecule #1: Cytochrome bc1 complex cytochrome c subunit
+Macromolecule #2: Cytochrome c oxidase subunit 1
+Macromolecule #3: Cytochrome bc1 complex Rieske iron-sulfur subunit
+Macromolecule #4: Cytochrome aa3 subunit 3
+Macromolecule #5: Cytochrome bc1 complex cytochrome b subunit
+Macromolecule #6: Putative conserved transmembrane protein
+Macromolecule #7: Superoxide dismutase [Cu-Zn]
+Macromolecule #8: Cytochrome aa3 subunit 2
+Macromolecule #9: Cytochrome c oxidase polypeptide 4
+Macromolecule #10: Cytochrome c oxidase subunit CtaJ
+Macromolecule #11: Uncharacterized protein MSMEG_4692/MSMEI_4575
+Macromolecule #12: LpqE protein
+Macromolecule #13: HEME C
+Macromolecule #14: (2R)-2-(hexadecanoyloxy)-3-{[(S)-hydroxy{[(1R,2R,3R,4R,5R,6S)-2,3...
+Macromolecule #15: MENAQUINONE-9
+Macromolecule #16: COPPER (II) ION
+Macromolecule #17: HEME-A
+Macromolecule #18: CARDIOLIPIN
+Macromolecule #19: FE2/S2 (INORGANIC) CLUSTER
+Macromolecule #20: (2R)-3-(((2-aminoethoxy)(hydroxy)phosphoryl)oxy)-2-(palmitoyloxy)...
+Macromolecule #21: PROTOPORPHYRIN IX CONTAINING FE
+Macromolecule #22: PALMITIC ACID
+Macromolecule #23: (2S)-1-(hexadecanoyloxy)propan-2-yl (10S)-10-methyloctadecanoate
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Average electron dose: 43.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 150885 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-7rh6: |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)