+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21927 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
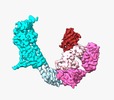
| Title | Structure of human Frizzled5 by fiducial-assisted cryo-EM | ||||||||||||
 Map data Map data | Sharpen map of the Fzd5-BRIL/Fab/Nb complex. | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology informationregulation of chorionic trophoblast cell proliferation / Spemann organizer formation / cellular response to molecule of bacterial origin / syncytiotrophoblast cell differentiation involved in labyrinthine layer development / embryonic camera-type eye morphogenesis / chorionic trophoblast cell differentiation / glandular epithelial cell maturation / Signaling by RNF43 mutants / post-embryonic camera-type eye development / apoptotic process involved in morphogenesis ...regulation of chorionic trophoblast cell proliferation / Spemann organizer formation / cellular response to molecule of bacterial origin / syncytiotrophoblast cell differentiation involved in labyrinthine layer development / embryonic camera-type eye morphogenesis / chorionic trophoblast cell differentiation / glandular epithelial cell maturation / Signaling by RNF43 mutants / post-embryonic camera-type eye development / apoptotic process involved in morphogenesis / WNT5A-dependent internalization of FZD2, FZD5 and ROR2 / anterior/posterior axis specification, embryo / embryonic axis specification / non-canonical Wnt signaling pathway / intestinal epithelial cell maturation / regulation of mitophagy / Wnt receptor activity / branching involved in labyrinthine layer morphogenesis / Wnt-protein binding / Class B/2 (Secretin family receptors) / regulation of bicellular tight junction assembly / Disassembly of the destruction complex and recruitment of AXIN to the membrane / positive regulation of protein targeting to mitochondrion / labyrinthine layer blood vessel development / bicellular tight junction / canonical Wnt signaling pathway / synapse assembly / Regulation of FZD by ubiquitination / positive regulation of interleukin-1 beta production / Asymmetric localization of PCP proteins / G protein-coupled receptor activity / electron transport chain / clathrin-coated endocytic vesicle membrane / neuron differentiation / positive regulation of T cell cytokine production / positive regulation of tumor necrosis factor production / positive regulation of type II interferon production / Ca2+ pathway / amyloid-beta binding / early endosome membrane / T cell differentiation in thymus / perikaryon / angiogenesis / periplasmic space / electron transfer activity / iron ion binding / negative regulation of cell population proliferation / Golgi membrane / axon / lipid binding / ubiquitin protein ligase binding / synapse / dendrite / heme binding / protein-containing complex binding / protein kinase binding / perinuclear region of cytoplasm / cell surface / positive regulation of transcription by RNA polymerase II / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||||||||
 Authors Authors | Tsutsumi N / Jude KM / Gati C / Garcia KC | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Structure of human Frizzled5 by fiducial-assisted cryo-EM supports a heterodimeric mechanism of canonical Wnt signaling. Authors: Naotaka Tsutsumi / Somnath Mukherjee / Deepa Waghray / Claudia Y Janda / Kevin M Jude / Yi Miao / John S Burg / Nanda Gowtham Aduri / Anthony A Kossiakoff / Cornelius Gati / K Christopher Garcia /   Abstract: Frizzleds (Fzd) are the primary receptors for Wnt morphogens, which are essential regulators of stem cell biology, yet the structural basis of Wnt signaling through Fzd remains poorly understood. ...Frizzleds (Fzd) are the primary receptors for Wnt morphogens, which are essential regulators of stem cell biology, yet the structural basis of Wnt signaling through Fzd remains poorly understood. Here we report the structure of an unliganded human Fzd5 determined by single-particle cryo-EM at 3.7 Å resolution, with the aid of an antibody chaperone acting as a fiducial marker. We also analyzed the topology of low-resolution XWnt8/Fzd5 complex particles, which revealed extreme flexibility between the Wnt/Fzd-CRD and the Fzd-TM regions. Analysis of Wnt/β-catenin signaling in response to Wnt3a versus a 'surrogate agonist' that cross-links Fzd to LRP6, revealed identical structure-activity relationships. Thus, canonical Wnt/β-catenin signaling appears to be principally reliant on ligand-induced Fzd/LRP6 heterodimerization, versus the allosteric mechanisms seen in structurally analogous class A G protein-coupled receptors, and Smoothened. These findings deepen our mechanistic understanding of Wnt signal transduction, and have implications for harnessing Wnt agonism in regenerative medicine. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21927.map.gz emd_21927.map.gz | 118 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21927-v30.xml emd-21927-v30.xml emd-21927.xml emd-21927.xml | 22.7 KB 22.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21927.png emd_21927.png | 134 KB | ||
| Others |  emd_21927_additional.map.gz emd_21927_additional.map.gz | 62.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21927 http://ftp.pdbj.org/pub/emdb/structures/EMD-21927 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21927 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21927 | HTTPS FTP |
-Validation report
| Summary document |  emd_21927_validation.pdf.gz emd_21927_validation.pdf.gz | 433.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21927_full_validation.pdf.gz emd_21927_full_validation.pdf.gz | 432.8 KB | Display | |
| Data in XML |  emd_21927_validation.xml.gz emd_21927_validation.xml.gz | 6.5 KB | Display | |
| Data in CIF |  emd_21927_validation.cif.gz emd_21927_validation.cif.gz | 7.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21927 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21927 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21927 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21927 | HTTPS FTP |
-Related structure data
| Related structure data |  6ww2MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10507 (Title: Structure of human Frizzled5 by fiducial-assisted cryo-EM EMPIAR-10507 (Title: Structure of human Frizzled5 by fiducial-assisted cryo-EMData size: 6.0 TB Data #1: Unaligned dark-subtracted TIFF movies with a gain reference for the 3D reconstruction of EMD-21927. [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21927.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21927.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpen map of the Fzd5-BRIL/Fab/Nb complex. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.078 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Unsharpen map of the Fzd5-BRIL/Fab/Nb complex.
| File | emd_21927_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpen map of the Fzd5-BRIL/Fab/Nb complex. | ||||||||||||
| Projections & Slices |
| ||||||||||||
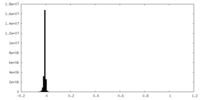
| Density Histograms |
- Sample components
Sample components
-Entire : Fzd5-BRIL/Fab/Nb complex
| Entire | Name: Fzd5-BRIL/Fab/Nb complex |
|---|---|
| Components |
|
-Supramolecule #1: Fzd5-BRIL/Fab/Nb complex
| Supramolecule | Name: Fzd5-BRIL/Fab/Nb complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Location in cell: Membrane Homo sapiens (human) / Location in cell: Membrane |
| Molecular weight | Theoretical: 100 KDa |
-Supramolecule #2: anti-BRIL Fab
| Supramolecule | Name: anti-BRIL Fab / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1, #3 |
|---|---|
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: HEK293S GnTi- Homo sapiens (human) / Recombinant strain: HEK293S GnTi- |
-Supramolecule #3: anti-Fab Nanobody
| Supramolecule | Name: anti-Fab Nanobody / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Recombinant expression | Organism: synthetic construct (others) |
-Supramolecule #4: Frizzled-5
| Supramolecule | Name: Frizzled-5 / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: anti-BRIL Fab Heavy chain
| Macromolecule | Name: anti-BRIL Fab Heavy chain / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.321084 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NVVDFSLHWV RQAPGKGLEW VAYISSSSGS TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARWGYWPGEP WWKAFDYWGQ GTLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW ...String: EISEVQLVES GGGLVQPGGS LRLSCAASGF NVVDFSLHWV RQAPGKGLEW VAYISSSSGS TSYADSVKGR FTISADTSKN TAYLQMNSL RAEDTAVYYC ARWGYWPGEP WWKAFDYWGQ GTLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW NSGALTSGVH TFPAVLQSSG LYSLSSVVTV PSSSLGTQTY ICNVNHKPSN TKVDKKVEPK S |
-Macromolecule #2: anti-Fab Nanobody
| Macromolecule | Name: anti-Fab Nanobody / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 13.390644 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSQVQLQESG GGLVQPGGSL RLSCAASGRT ISRYAMSWFR QAPGKEREFV AVARRSGDGA FYADSVQGRF TVSRDDAKNT VYLQMNSLK PEDTAVYYCA IDSDTFYSGS YDYWGQGTQV TVSS |
-Macromolecule #3: anti-BRIL Fab Light chain
| Macromolecule | Name: anti-BRIL Fab Light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.353947 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQYLYYSLVT FGQGTKVEIK RTVAAPSVFI FPPSDSQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG N SQESVTEQ ...String: SDIQMTQSPS SLSASVGDRV TITCRASQSV SSAVAWYQQK PGKAPKLLIY SASSLYSGVP SRFSGSRSGT DFTLTISSLQ PEDFATYYC QQYLYYSLVT FGQGTKVEIK RTVAAPSVFI FPPSDSQLKS GTASVVCLLN NFYPREAKVQ WKVDNALQSG N SQESVTEQ DSKDSTYSLS STLTLSKADY EKHKVYACEV THQGLSSPVT KSFNRG |
-Macromolecule #4: Frizzled-5,Soluble cytochrome b562
| Macromolecule | Name: Frizzled-5,Soluble cytochrome b562 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 74.75418 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DYKDDDDASK APVCQEITVP MCRGIGYNLT HMPNQFNHDT QDEAGLEVHQ FWPLVEIQCS PDLRFFLCSM YTPICLPDYH KPLPPCRSV CERAKAGCSP LMRQYGFAWP ERMSCDRLPV LGRDAEVLCM DYNRSEATTA PPRPFPAKPT LPGPPGAPAS G GECPAGGP ...String: DYKDDDDASK APVCQEITVP MCRGIGYNLT HMPNQFNHDT QDEAGLEVHQ FWPLVEIQCS PDLRFFLCSM YTPICLPDYH KPLPPCRSV CERAKAGCSP LMRQYGFAWP ERMSCDRLPV LGRDAEVLCM DYNRSEATTA PPRPFPAKPT LPGPPGAPAS G GECPAGGP FVCKCREPFV PILKESHPLY NKVRTGQVPN CAVPCYQPSF SADERTFATF WIGLWSVLCF ISTSTTVATF LI DMERFRY PERPIIFLSA CYLCVSLGFL VRLVVGHASV ACSREHNHIH YETTGPALCT IVFLLVYFFG MASSIWWVIL SLT WFLAAG MKWGNEAIAG YAQYFHLAAW LIPSVKSITA LALSSVDGDP VAGICYVGNQ NLNSLRGFVL GPLVLYLLVG TLFL LAGFV SLFRARRQLA DLEDNWETLN DNLKVIEKAD NAAQVKDALT KMRAAALDAQ KATPPKLEDK SPDSPEMKDF RHGFD ILVG QIDDALKLAN EGKVKEAQAA AEQLKTTRNA YIQKYLERAR STLDKLEKLM IRIGIFTLLY TVPASIVVAC YLYEQH YRE SWEAALTCAC PGHDTGQPRA KPEYWVLMLK YFMCLVVGIT SGVWIWSGKT VESWRRFTSR CCCRPRRGHK AAALEVL FQ GPGAAEDQVD PRLIDGKHHH HHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.2 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 281 K / Instrument: LEICA EM GP / Details: 5s blotting. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 10898 / Average exposure time: 0.1225 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated magnification: 81000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: -2.0 µm / Nominal defocus min: -0.8 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6ww2: |
 Movie
Movie Controller
Controller



















 Z
Z Y
Y X
X