+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21251 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of apo-BBSome | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationestablishment of anatomical structure orientation / BBSome-mediated cargo-targeting to cilium / receptor localization to non-motile cilium / multi-ciliated epithelial cell differentiation / BBSome / camera-type eye photoreceptor cell differentiation / renal tubule development / smoothened binding / establishment of planar polarity / inner ear receptor cell stereocilium organization ...establishment of anatomical structure orientation / BBSome-mediated cargo-targeting to cilium / receptor localization to non-motile cilium / multi-ciliated epithelial cell differentiation / BBSome / camera-type eye photoreceptor cell differentiation / renal tubule development / smoothened binding / establishment of planar polarity / inner ear receptor cell stereocilium organization / photoreceptor connecting cilium / patched binding / olfactory bulb development / establishment of epithelial cell apical/basal polarity / protein localization to cilium / phosphatidylinositol-3-phosphate binding / regulation of stress fiber assembly / non-motile cilium assembly / non-motile cilium / motile cilium / sensory perception / centrosome cycle / eating behavior / ciliary membrane / erythrocyte homeostasis / pericentriolar material / ciliary transition zone / B cell homeostasis / cilium assembly / axoneme / fat cell differentiation / axon guidance / protein localization to plasma membrane / multicellular organism growth / fibrillar center / centriolar satellite / Wnt signaling pathway / sensory perception of smell / intracellular protein localization / regulation of protein localization / protein transport / gene expression / RNA polymerase II-specific DNA-binding transcription factor binding / protein-macromolecule adaptor activity / neuron projection / cilium / ciliary basal body / centrosome / membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.44 Å | |||||||||
 Authors Authors | Yang S / Walz T | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Near-atomic structures of the BBSome reveal the basis for BBSome activation and binding to GPCR cargoes. Authors: Shuang Yang / Kriti Bahl / Hui-Ting Chou / Jonathan Woodsmith / Ulrich Stelzl / Thomas Walz / Maxence V Nachury /   Abstract: Dynamic trafficking of G protein-coupled receptors (GPCRs) out of cilia is mediated by the BBSome. In concert with its membrane recruitment factor, the small GTPase ARL6/BBS3, the BBSome ferries ...Dynamic trafficking of G protein-coupled receptors (GPCRs) out of cilia is mediated by the BBSome. In concert with its membrane recruitment factor, the small GTPase ARL6/BBS3, the BBSome ferries GPCRs across the transition zone, a diffusion barrier at the base of cilia. Here, we present the near-atomic structures of the BBSome by itself and in complex with ARL6, and we describe the changes in BBSome conformation induced by ARL6 binding. Modeling the interactions of the BBSome with membranes and the GPCR Smoothened (SMO) reveals that SMO, and likely also other GPCR cargoes, must release their amphipathic helix 8 from the membrane to be recognized by the BBSome. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21251.map.gz emd_21251.map.gz | 72.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21251-v30.xml emd-21251-v30.xml emd-21251.xml emd-21251.xml | 21.3 KB 21.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_21251.png emd_21251.png | 31.6 KB | ||
| Filedesc metadata |  emd-21251.cif.gz emd-21251.cif.gz | 8.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21251 http://ftp.pdbj.org/pub/emdb/structures/EMD-21251 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21251 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21251 | HTTPS FTP |
-Related structure data
| Related structure data |  6vnwMC  6voaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21251.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21251.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
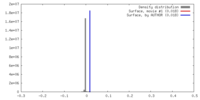
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : BBSome complex
| Entire | Name: BBSome complex |
|---|---|
| Components |
|
-Supramolecule #1: BBSome complex
| Supramolecule | Name: BBSome complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 500 KDa |
-Macromolecule #1: Bardet-Biedl syndrome 18 protein
| Macromolecule | Name: Bardet-Biedl syndrome 18 protein / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 8.070502 KDa |
| Sequence | String: MAETKSMFRE VLPKQGQLYV EDITTMVLCK PKLLPLKSLT LEKLEKMQQA AQDTIHQQEM TEKEQKITH UniProtKB: BBSome interacting protein 1 |
-Macromolecule #2: Bardet-Biedl syndrome 2 protein homolog
| Macromolecule | Name: Bardet-Biedl syndrome 2 protein homolog / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 79.911484 KDa |
| Sequence | String: MLQPVFTLKL RHKISPRMVA VGRYDGTHPC LAAATQAGKV FIHNPHSRSQ HLGAPRVLQS PLESDVSLLN INQTVSCLTA GVLNPELGY DALLVGTQTN LLAYDVYNNS DLFYREVADG ASAIVLGTLG DITSPLAIIG GNCALQGFNH EGNDLFWTVT G DNVHSLAL ...String: MLQPVFTLKL RHKISPRMVA VGRYDGTHPC LAAATQAGKV FIHNPHSRSQ HLGAPRVLQS PLESDVSLLN INQTVSCLTA GVLNPELGY DALLVGTQTN LLAYDVYNNS DLFYREVADG ASAIVLGTLG DITSPLAIIG GNCALQGFNH EGNDLFWTVT G DNVHSLAL CDFDGDGKKE LLVGSEDFDI RVFKEDEIVA EMSETEIITS LCPMYGSRFG YALSNGTVGV YDKTARYWRI KS KNQAMSI HAFDLNSDGV CELITGWSNG KVDARSDRTG EVIFKDNFSS AIAGVVEGDY RMEGCQQLIC CSVDGEIRGY LPG TAEMRG NLMDISVEQD LIRELSQKKQ NLLLELRNYE ENAKAELSSP LNEADGHRGV IPANTKHHTA LSVSLGSEAQ AAHA ELCIS TSNDTIIRAV LIFAEGVFAG ESHVVHPSVH HLSSSVRIPI TPPKDIPVDL HLKTFVGYRS STQFHVFELT RQLPR FSMY ALTSPDPASE PLSYVNFIIA ERAQRVVMWL NQNFLLPEDT NIQNAPFQVC FTSLRNGGQL YIKIKLSGEI TVNTDD IDL AGDIIQSMAS FFAIEDLQVE ADFPVYFEEL RKVLVKVDEY HSVHQKLSAD MADNSNLIRS LLVQAEDARL MRDMKTM KN RYKELYDLNK DLLNGYKIRC NNHTELLGSL KAVNQAIQRA GHLRVGKPKN QVITACRDAI RSNNINMLFR IMRVGTAS S UniProtKB: Bardet-Biedl syndrome 2 protein homolog |
-Macromolecule #3: Bardet-Biedl syndrome 4 protein homolog
| Macromolecule | Name: Bardet-Biedl syndrome 4 protein homolog / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 58.289133 KDa |
| Sequence | String: MAEEKLSART QLPVSAESQK PVLKKAPEFP ILEKQNWLIH LYYIQKDYEA CKAVIKEQLQ ETHGLCEYAI YVQALIFRLE GNIQESLRL FQMCAFLSPQ CADNLKQVAR SLFLLGKHKA AIEVYNEAAK LNQKDWEICH NLGVCYIYLK QFDKAQDQLH N ALHLNRHD ...String: MAEEKLSART QLPVSAESQK PVLKKAPEFP ILEKQNWLIH LYYIQKDYEA CKAVIKEQLQ ETHGLCEYAI YVQALIFRLE GNIQESLRL FQMCAFLSPQ CADNLKQVAR SLFLLGKHKA AIEVYNEAAK LNQKDWEICH NLGVCYIYLK QFDKAQDQLH N ALHLNRHD LTYIMLGKIF LLKGDLDKAI EIYKKAVEFS PENTELLTTL GLLYLQLGIY QKAFEHLGNT LTYDPTNYKA IL AAGSMMQ THGDFDVALT KYKVVACAVI ESPPLWNNIG MCFFGKKKYV AAISCLKRAN YLAPLDWKIL YNLGLVHLTM QQY ASAFHF LSAAINFQPK MGELYMLLAV ALTNLEDSEN AKRAYEEAVR LDKCNPLVNL NYAVLLYNQG EKRDALAQYQ EMEK KVNLL KYSSSLEFDP EMVEVAQKLG AALQVGEALV WTKPVKDPKS KHQTASTSKA AGFQQPLGSN QALGQAMSSA ATCRK LSSG AGGTSQLTKP PSLPLEPEPT VEAQPTEASA QTREK UniProtKB: BBSome complex member BBS4 |
-Macromolecule #4: Bardet-Biedl syndrome 5 protein homolog
| Macromolecule | Name: Bardet-Biedl syndrome 5 protein homolog / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.880984 KDa |
| Sequence | String: MSVLDALWED RDVRFDVSSQ QMKTRPGEVL IDCLDSVEDT KGNNGDRGRL LVTNLRIVWH SLALPRVNLS IGYNCILNIT TRTANSKLR GQTEALYVLT KCNSTRFEFI FTNLVPGSPR LYTSLIAVHR AYETSKMYRD FKLRSALIQN KQLRLLPQEN V YNKINGVW ...String: MSVLDALWED RDVRFDVSSQ QMKTRPGEVL IDCLDSVEDT KGNNGDRGRL LVTNLRIVWH SLALPRVNLS IGYNCILNIT TRTANSKLR GQTEALYVLT KCNSTRFEFI FTNLVPGSPR LYTSLIAVHR AYETSKMYRD FKLRSALIQN KQLRLLPQEN V YNKINGVW NLSSDQGNLG TFFITNVRIV WHANMNDSFN VSIPYLQIRS VKIRDSKFGL ALVIESSQQS GGYVLGFKID PV EKLQESV KEINSLHKVY SANPIFGVDY EMEEKPQPLE ALTVKQIQDD VEIDSDDHTD AFVAYFADGN KQQDREPVFS EEL GLAIEK LKDGFTLQGL WEVMN UniProtKB: Bardet-Biedl syndrome 5 protein homolog |
-Macromolecule #5: Bardet-Biedl syndrome 7 protein homolog
| Macromolecule | Name: Bardet-Biedl syndrome 7 protein homolog / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 80.471375 KDa |
| Sequence | String: MDLNLNRADY LQVGVTSQKT MKLLPASKHR ATQKVVVGDH DGIVMCFGMK KGEAVTVFKT LPGQKIARLE LGGALNTPQE KIFIAAGSE IRGFTKRGKQ FLSFETNLTE SIKAMHISGS DLFLSASYIY NHYCDCKDQH YYLSGDKIND VICLPVERLL R EVPVLACQ ...String: MDLNLNRADY LQVGVTSQKT MKLLPASKHR ATQKVVVGDH DGIVMCFGMK KGEAVTVFKT LPGQKIARLE LGGALNTPQE KIFIAAGSE IRGFTKRGKQ FLSFETNLTE SIKAMHISGS DLFLSASYIY NHYCDCKDQH YYLSGDKIND VICLPVERLL R EVPVLACQ DRVLRVLQGS DVTYEIEVPG PPTVLALHNG NGGDSGEDLL FGTSDGKLGL IQITTSKPIH KWEIRNEKKR GG ILCVDSF DIVGDGVKDL LVGRDDGMVE VYGFDNANEP VLRFDHTLSE SVTSIQGGCV GKDGYDEIVV STYSGWITGL TTE PVHKES GPGEELKFNQ EMQNKISSLR SELEQLQYKV LQEREKYQQS SQSSKAKSAV PSFSVNDKFT LNKDDASYSL ILEV QTAID NVLIQSDVPI DLLDVDKNSA VVSFSSCDSE SNDNFLLATY RCQANTTRLE LKIRSIEGQY GTLQAYVTPR IQPKT CQVR QYHIKPLSLH QRTHFIDHDR PMNTLTLTGQ FSFSELHSWV VFCMPEVPEK PPAGECVTFY FQNTFLDTQL ESTYRK GEG VFKSDNISTI SILKDVLSKE ATKRKINLNI SYEINEVSVK HTLKLIHPKL EYQLLLAKKV QLIDALKELQ VHEGNTN FL IPEYRCILEE ADHLQEEYKK QPAHLERLYG MITDLFIDKF KFKGTNVKTK VPLLLEILDS YDQNALIAFF DAA UniProtKB: Bardet-Biedl syndrome 7 protein homolog |
-Macromolecule #6: Tetratricopeptide repeat domain 8
| Macromolecule | Name: Tetratricopeptide repeat domain 8 / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 56.686406 KDa |
| Sequence | String: MEPLLLAWSY FRRRRFQLCA DLCTQMLEKS PCDQAAWILK ARALTEMVYV DEIDVDEEGI AEMILDENAI AQVPRPGTSL KLPGTNQTG GPSPAVRPVT QAGRPITGFL RPSTQSGRPG TIEQAIKTPR TAYTARPIAS SSGRFVRLGT ASMLTSPDGP F INLSRLNL ...String: MEPLLLAWSY FRRRRFQLCA DLCTQMLEKS PCDQAAWILK ARALTEMVYV DEIDVDEEGI AEMILDENAI AQVPRPGTSL KLPGTNQTG GPSPAVRPVT QAGRPITGFL RPSTQSGRPG TIEQAIKTPR TAYTARPIAS SSGRFVRLGT ASMLTSPDGP F INLSRLNL AKYAQKPKLA KALFEYIFHH ENDVKTALDL AALSTEHSQY KDWWWKVQIG KCYYRLGLYR EAEKQFKSAL KQ QEMVDTF LYLAKVYISL DQPLTALNLF KQGLDKFPGE VTLLCGIARI YEEMNNISSA TEYYKEVLKQ DNTHVEAIAC IGS NHFYTD QPEVALRFYR RLLQMGVYNC QLFNNLGLCC FYAQQYDMTL TSFERALSLA ENEEEVADVW YNLGHVAVGT GDTN LAHQC FRLALVSNNQ HAEAYNNLAV LEMRRGHVEQ AKALLQTASS LAPHMYEPHF NFATISDKIG DLQRSYAAAK KSEAA FPDH VDTQHLIKQL EQHFAML UniProtKB: Tetratricopeptide repeat domain 8 |
-Macromolecule #7: Bardet-Biedl syndrome 9
| Macromolecule | Name: Bardet-Biedl syndrome 9 / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 99.230914 KDa |
| Sequence | String: MSLFKARDWW STVLGDKEEF DQGCLCLADV DNTGNGQDKI IVGSFMGYLR IFNPHPVKTG DGAQAEDLLL EVHLRDPILQ VEVGKFVSG TEMLHLAVLH SRKLCVYSVS GTLGNVEHGN QYQIKLMYEH NLQRTACNMT YGSFGGVKGR DLICIQSVDG M LMVFEQES ...String: MSLFKARDWW STVLGDKEEF DQGCLCLADV DNTGNGQDKI IVGSFMGYLR IFNPHPVKTG DGAQAEDLLL EVHLRDPILQ VEVGKFVSG TEMLHLAVLH SRKLCVYSVS GTLGNVEHGN QYQIKLMYEH NLQRTACNMT YGSFGGVKGR DLICIQSVDG M LMVFEQES YAFGRFLPGS LLPGPLAYSS RTDSFITVSS CHQVESYKYQ VLAFATDADK RQETEQQKHG SGKRLVVDWT LN IGEQAID ICIVSFIQSA SSVFVLGERN FFCLKDNGQI QFMKKLDYSP SCFLPYCSVS EGTINTLIGN HNNMLHIYQD VTL KWATQL PHVPVAVRVG CLHDLKGVIV TLSDDGHLQC SYLGTDPSLF QAPKVESREL NYDELDMELK ELQKVIKNVN KSQD VWPLT EREDDLKVSA MVSPNFDSVS QATDVEVGAD LVPSVTVKVT LKNRVALQKI KLSIYVQPPL VLTGDQFTFE FMAPE MTRT VGFSVYLKGS YSPPELEGNA VVSYSRPTER NPDGIPRVSQ CKFRLPLKLV CLPGQPSKTA SHKLTIDTNK SPVSLL SLF PGFAKQSEDD QVNVMGFRFL GGSQVTLLAS KTSQRYRIQS EQFEDLWLIT NELIIRLQEY FEKQGIKDFT CSFSGSV PL EEYFELIDHH FELRINGEKL EELLSERAVQ FRAIQRRLLT RFKDKTPAPL QHLDTLLDGT YKQVIALADA VEENQDNL F QSFTRLKSAT HLVILLIGLW QKLSADQIAI LEAAFLPLQQ DTQELGWEET VDAALSHLLK TCLSKSSKEQ ALNLNSQLG IPKDTSQLKK HITLFCDRLA KGGRLCLSTD AAAPQTMVMP GGCATIPESD LEGRSIDQDS SELFTNHKHL MVETPVPEVS PLQGVTE UniProtKB: Bardet-Biedl syndrome 9 |
-Macromolecule #8: BBS1 domain-containing protein
| Macromolecule | Name: BBS1 domain-containing protein / type: protein_or_peptide / ID: 8 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 64.939141 KDa |
| Sequence | String: MAATSSSDSD GGKGESEANS KWLDSLSDSM ANIHTFSACL ALADFHGDGE YKLAMGDLGP DGRQPRLKVL KGHTLVSQKP LPDLPAAAV TFLMASHEPR TPALAIASGP CVYVYKNLKP YFKFSLPSLP TNPLEQDLWN QAKEDQIDPL TLKEMLEGIR E KAEVPLSV ...String: MAATSSSDSD GGKGESEANS KWLDSLSDSM ANIHTFSACL ALADFHGDGE YKLAMGDLGP DGRQPRLKVL KGHTLVSQKP LPDLPAAAV TFLMASHEPR TPALAIASGP CVYVYKNLKP YFKFSLPSLP TNPLEQDLWN QAKEDQIDPL TLKEMLEGIR E KAEVPLSV QSLRFLPLEL SEMEAFVNQH KSKSIRRQTV ITTMTTLKKN LADEDAVSCL VLGTENKELL VLDPEAFTIL AK MSLPSVP AFLEASGQFD VEFRLAAACR NGSIYILRRD SKRPKYCIEL GAQPVGLVGV HKVLVVGSNQ DSLHGFTYKG KRL WTVQMP AAILAMNLLE QHSRGLQAVM AALANEEVRI YHDKVLLNVI RTPEAVTSLC FGRYGREDNT LIMTTLGGGL IIKI LKRTA VFAEGGGEAG PPPSQAIKLN VPRKTRLYVD QTLREREAGT AMHRTFQADL YLLRLRAARA YVQALESSLS PVSLT AREP LKLHAVVQGL GPTFKLTLHL QNTSTARPIL GLVVCFLYNE VLYALPRAFF KVPLLVPGLN YPLETFVKSL SDKGIS DII KVLVLREGQS TPLLSAHINM PMSEGLAAD UniProtKB: Bardet-Biedl syndrome 1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. / Pretreatment - Atmosphere: AIR | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: Wait time 20s, blot force -2, blot time 3.5s. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 80.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP EMDB ID: |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 3.44 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 560777 |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)