[English] 日本語
 Yorodumi
Yorodumi- EMDB-17696: Structure of human 48S translation initiation complex in open cod... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of human 48S translation initiation complex in open codon scanning state (48S-1) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RIBOSOME / TRANSLATION / initiation / 48S / eIF / human / eukaryotic / factor / codon / scanning / open / reading | |||||||||
| Function / homology |  Function and homology information Function and homology informationmale germ cell proliferation / positive regulation of mRNA binding / regulation of translation in response to endoplasmic reticulum stress / translation initiation ternary complex / positive regulation of mRNA cis splicing, via spliceosome / glial limiting end-foot / response to kainic acid / HRI-mediated signaling / Cellular response to mitochondrial stress / viral translational termination-reinitiation ...male germ cell proliferation / positive regulation of mRNA binding / regulation of translation in response to endoplasmic reticulum stress / translation initiation ternary complex / positive regulation of mRNA cis splicing, via spliceosome / glial limiting end-foot / response to kainic acid / HRI-mediated signaling / Cellular response to mitochondrial stress / viral translational termination-reinitiation / response to manganese-induced endoplasmic reticulum stress / positive regulation of type B pancreatic cell apoptotic process / methionyl-initiator methionine tRNA binding / eukaryotic translation initiation factor 3 complex, eIF3e / negative regulation of translational initiation in response to stress / Response of EIF2AK1 (HRI) to heme deficiency / cap-dependent translational initiation / eukaryotic translation initiation factor 3 complex, eIF3m / Recycling of eIF2:GDP / PERK-mediated unfolded protein response / translation reinitiation / selenocysteine metabolic process / PERK regulates gene expression / eukaryotic translation initiation factor 2 complex / IRES-dependent viral translational initiation / regulation of translational initiation in response to stress / multi-eIF complex / formation of cytoplasmic translation initiation complex / eukaryotic translation initiation factor 3 complex / selenocysteine incorporation / selenocysteine insertion sequence binding / protein-synthesizing GTPase / eukaryotic 43S preinitiation complex / cytoplasmic translational initiation / translation factor activity, RNA binding / mRNA cap binding / formation of translation preinitiation complex / positive regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis / negative regulation of endoplasmic reticulum unfolded protein response / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair / eukaryotic 48S preinitiation complex / protein tyrosine kinase inhibitor activity / positive regulation of respiratory burst involved in inflammatory response / regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage / positive regulation of gastrulation / nucleolus organization / IRE1-RACK1-PP2A complex / : / positive regulation of endodeoxyribonuclease activity / positive regulation of Golgi to plasma membrane protein transport / TNFR1-mediated ceramide production / negative regulation of DNA repair / negative regulation of RNA splicing / metal-dependent deubiquitinase activity / negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide / oxidized purine DNA binding / supercoiled DNA binding / neural crest cell differentiation / NF-kappaB complex / ubiquitin-like protein conjugating enzyme binding / regulation of translational initiation / regulation of establishment of cell polarity / negative regulation of phagocytosis / positive regulation of ubiquitin-protein transferase activity / rRNA modification in the nucleus and cytosol / Formation of the ternary complex, and subsequently, the 43S complex / erythrocyte homeostasis / cytoplasmic side of rough endoplasmic reticulum membrane / laminin receptor activity / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / protein kinase A binding / negative regulation of ubiquitin protein ligase activity / Ribosomal scanning and start codon recognition / ion channel inhibitor activity / Translation initiation complex formation / pigmentation / mammalian oogenesis stage / positive regulation of mitochondrial depolarization / activation-induced cell death of T cells / positive regulation of T cell receptor signaling pathway / negative regulation of Wnt signaling pathway / fibroblast growth factor binding / positive regulation of activated T cell proliferation / iron-sulfur cluster binding / regulation of cell division / Protein hydroxylation / negative regulation of peptidyl-serine phosphorylation / BH3 domain binding / mTORC1-mediated signalling / SARS-CoV-1 modulates host translation machinery / monocyte chemotaxis / Peptide chain elongation / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / cysteine-type endopeptidase activator activity involved in apoptotic process / Selenocysteine synthesis / positive regulation of signal transduction by p53 class mediator / Formation of a pool of free 40S subunits Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
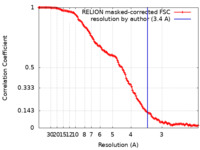
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Petrychenko V / Yi S-H / Liedtke D / Peng BZ / Rodnina MV / Fischer N | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nat.Struct.Mol.Biol. / Year: 2024 Journal: Nat.Struct.Mol.Biol. / Year: 2024Title: Structural basis for translational control by the human 48S initiation complex from codon scanning toward subunit joining Authors: Petrychenko V / Yi S-H / Liedtke D / Peng BZ / Rodnina MV / Fischer N | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_17696.map.gz emd_17696.map.gz | 275.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-17696-v30.xml emd-17696-v30.xml emd-17696.xml emd-17696.xml | 87.3 KB 87.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_17696_fsc.xml emd_17696_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_17696.png emd_17696.png | 146.1 KB | ||
| Masks |  emd_17696_msk_1.map emd_17696_msk_1.map | 178 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-17696.cif.gz emd-17696.cif.gz | 19.1 KB | ||
| Others |  emd_17696_additional_1.map.gz emd_17696_additional_1.map.gz emd_17696_additional_2.map.gz emd_17696_additional_2.map.gz emd_17696_additional_3.map.gz emd_17696_additional_3.map.gz emd_17696_additional_4.map.gz emd_17696_additional_4.map.gz emd_17696_half_map_1.map.gz emd_17696_half_map_1.map.gz emd_17696_half_map_2.map.gz emd_17696_half_map_2.map.gz | 162.2 MB 44.3 MB 5.1 MB 5.1 MB 140.6 MB 140.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-17696 http://ftp.pdbj.org/pub/emdb/structures/EMD-17696 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17696 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-17696 | HTTPS FTP |
-Related structure data
| Related structure data |  8pj1MC  17697  17698  17699  17700  17701  19128  8pj2C  8pj3C  8pj4C  8pj5C  8pj6C  8rg0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_17696.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_17696.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.967 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_17696_msk_1.map emd_17696_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Human 48S initiation complex 40S-eIF1-eIF1A-eIF2-eIF3-tRNA-Met-mRNA
+Supramolecule #1: Human 48S initiation complex 40S-eIF1-eIF1A-eIF2-eIF3-tRNA-Met-mRNA
+Macromolecule #1: Eukaryotic translation initiation factor 3 subunit B
+Macromolecule #2: Eukaryotic translation initiation factor 3 subunit I
+Macromolecule #3: Eukaryotic translation initiation factor 3 subunit K
+Macromolecule #4: Eukaryotic translation initiation factor 3 subunit F
+Macromolecule #5: Eukaryotic translation initiation factor 3 subunit L
+Macromolecule #6: Eukaryotic translation initiation factor 3 subunit M
+Macromolecule #8: Eukaryotic translation initiation factor 3 subunit H
+Macromolecule #9: 60S ribosomal protein L41
+Macromolecule #11: 40S ribosomal protein S11
+Macromolecule #12: 40S ribosomal protein S4, X isoform
+Macromolecule #13: 40S ribosomal protein S9
+Macromolecule #14: 40S ribosomal protein S23
+Macromolecule #15: 40S ribosomal protein S30
+Macromolecule #16: 40S ribosomal protein S7
+Macromolecule #17: 40S ribosomal protein S27
+Macromolecule #18: 40S ribosomal protein S13
+Macromolecule #19: 40S ribosomal protein S15a
+Macromolecule #20: 40S ribosomal protein S21
+Macromolecule #21: 40S ribosomal protein S2
+Macromolecule #22: 40S ribosomal protein S17
+Macromolecule #23: 40S ribosomal protein SA
+Macromolecule #24: 40S ribosomal protein S3a
+Macromolecule #25: 40S ribosomal protein S14
+Macromolecule #26: 40S ribosomal protein S26
+Macromolecule #27: 40S ribosomal protein S8
+Macromolecule #28: 40S ribosomal protein S6
+Macromolecule #29: 40S ribosomal protein S24
+Macromolecule #30: 40S ribosomal protein S5
+Macromolecule #31: 40S ribosomal protein S16
+Macromolecule #32: 40S ribosomal protein S3
+Macromolecule #33: 40S ribosomal protein S10
+Macromolecule #34: 40S ribosomal protein S15
+Macromolecule #35: Receptor of activated protein C kinase 1
+Macromolecule #36: 40S ribosomal protein S19
+Macromolecule #37: 40S ribosomal protein S25
+Macromolecule #38: 40S ribosomal protein S18
+Macromolecule #39: 40S ribosomal protein S20
+Macromolecule #40: 40S ribosomal protein S29
+Macromolecule #41: Ubiquitin
+Macromolecule #42: 40S ribosomal protein S12
+Macromolecule #43: 40S ribosomal protein S28
+Macromolecule #44: Eukaryotic translation initiation factor 3 subunit G
+Macromolecule #45: Eukaryotic translation initiation factor 1
+Macromolecule #46: Eukaryotic translation initiation factor 1A, X-chromosomal
+Macromolecule #47: Eukaryotic translation initiation factor 2 subunit 1
+Macromolecule #48: Eukaryotic translation initiation factor 2 subunit 2
+Macromolecule #49: Eukaryotic translation initiation factor 2 subunit 3
+Macromolecule #50: Eukaryotic translation initiation factor 3 subunit A
+Macromolecule #51: Eukaryotic translation initiation factor 3 subunit E
+Macromolecule #53: Eukaryotic translation initiation factor 3 subunit D
+Macromolecule #54: Eukaryotic translation initiation factor 3 subunit C
+Macromolecule #7: mRNA
+Macromolecule #10: 18S rRNA
+Macromolecule #52: Initiator Met-tRNA-i
+Macromolecule #55: MAGNESIUM ION
+Macromolecule #56: ZINC ION
+Macromolecule #57: GUANOSINE-5'-TRIPHOSPHATE
+Macromolecule #58: METHIONINE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: HOMEMADE PLUNGER / Details: Manual blotting & plunge-freezing. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: Electron-optical aberrations were corrected using a CETCOR Cs-corrector (CEOS, Heidelberg) aligned with the CETCORPLUS 4.6.9 software package (CEOS, Heidelberg). |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Average exposure time: 1.5 sec. / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.2 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-8pj1: |
 Movie
Movie Controller
Controller









































 Z
Z Y
Y X
X