+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-13065 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of ALC1/CHD1L bound to a PARylated nucleosome | |||||||||||||||
 マップデータ マップデータ | Main map | |||||||||||||||
 試料 試料 |
| |||||||||||||||
 キーワード キーワード | ALC1 / CHD1L / chromatin remodeler / DNA damage response / nucleosome / poly(ADP-ribose) / DNA BINDING PROTEIN | |||||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報poly-ADP-D-ribose modification-dependent protein binding / ATP-dependent chromatin remodeler activity / site of DNA damage / nucleosome binding / DNA helicase activity / histone reader activity / 加水分解酵素; 酸無水物に作用; 酸無水物に作用・細胞または細胞小器官の運動に関与 / Dual Incision in GG-NER / Formation of Incision Complex in GG-NER / structural constituent of chromatin ...poly-ADP-D-ribose modification-dependent protein binding / ATP-dependent chromatin remodeler activity / site of DNA damage / nucleosome binding / DNA helicase activity / histone reader activity / 加水分解酵素; 酸無水物に作用; 酸無水物に作用・細胞または細胞小器官の運動に関与 / Dual Incision in GG-NER / Formation of Incision Complex in GG-NER / structural constituent of chromatin / nucleosome / nucleosome assembly / site of double-strand break / chromatin remodeling / protein heterodimerization activity / DNA repair / nucleotide binding / DNA damage response / ATP hydrolysis activity / DNA binding / nucleoplasm / ATP binding / nucleus / plasma membrane / cytosol 類似検索 - 分子機能 | |||||||||||||||
| 生物種 |  Homo sapiens (ヒト) / synthetic construct (人工物) Homo sapiens (ヒト) / synthetic construct (人工物) | |||||||||||||||
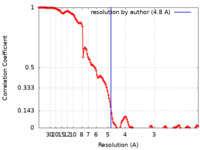
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 4.8 Å | |||||||||||||||
 データ登録者 データ登録者 | Bacic L / Gaullier G | |||||||||||||||
| 資金援助 |  スウェーデン, 4件 スウェーデン, 4件
| |||||||||||||||
 引用 引用 |  ジャーナル: Elife / 年: 2021 ジャーナル: Elife / 年: 2021タイトル: Structure and dynamics of the chromatin remodeler ALC1 bound to a PARylated nucleosome. 著者: Luka Bacic / Guillaume Gaullier / Anton Sabantsev / Laura C Lehmann / Klaus Brackmann / Despoina Dimakou / Mario Halic / Graeme Hewitt / Simon J Boulton / Sebastian Deindl /    要旨: The chromatin remodeler ALC1 is recruited to and activated by DNA damage-induced poly(ADP-ribose) (PAR) chains deposited by PARP1/PARP2/HPF1 upon detection of DNA lesions. ALC1 has emerged as a ...The chromatin remodeler ALC1 is recruited to and activated by DNA damage-induced poly(ADP-ribose) (PAR) chains deposited by PARP1/PARP2/HPF1 upon detection of DNA lesions. ALC1 has emerged as a candidate drug target for cancer therapy as its loss confers synthetic lethality in homologous recombination-deficient cells. However, structure-based drug design and molecular analysis of ALC1 have been hindered by the requirement for PARylation and the highly heterogeneous nature of this post-translational modification. Here, we reconstituted an ALC1 and PARylated nucleosome complex modified in vitro using PARP2 and HPF1. This complex was amenable to cryo-EM structure determination without cross-linking, which enabled visualization of several intermediate states of ALC1 from the recognition of the PARylated nucleosome to the tight binding and activation of the remodeler. Functional biochemical assays with PARylated nucleosomes highlight the importance of nucleosomal epitopes for productive remodeling and suggest that ALC1 preferentially slides nucleosomes away from DNA breaks. | |||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_13065.map.gz emd_13065.map.gz | 121.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-13065-v30.xml emd-13065-v30.xml emd-13065.xml emd-13065.xml | 27.3 KB 27.3 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_13065_fsc.xml emd_13065_fsc.xml | 13.9 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_13065.png emd_13065.png | 79.2 KB | ||
| マスクデータ |  emd_13065_msk_1.map emd_13065_msk_1.map | 244.1 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-13065.cif.gz emd-13065.cif.gz | 7.8 KB | ||
| その他 |  emd_13065_half_map_1.map.gz emd_13065_half_map_1.map.gz emd_13065_half_map_2.map.gz emd_13065_half_map_2.map.gz | 226.8 MB 226.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13065 http://ftp.pdbj.org/pub/emdb/structures/EMD-13065 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13065 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13065 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_13065_validation.pdf.gz emd_13065_validation.pdf.gz | 1.1 MB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_13065_full_validation.pdf.gz emd_13065_full_validation.pdf.gz | 1.1 MB | 表示 | |
| XML形式データ |  emd_13065_validation.xml.gz emd_13065_validation.xml.gz | 22.1 KB | 表示 | |
| CIF形式データ |  emd_13065_validation.cif.gz emd_13065_validation.cif.gz | 28.5 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13065 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13065 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13065 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-13065 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_13065.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_13065.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Main map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.0752 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
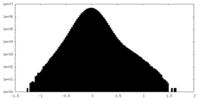
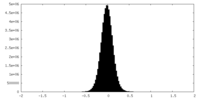
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-マスク #1
| ファイル |  emd_13065_msk_1.map emd_13065_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
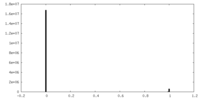
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Half map 2
| ファイル | emd_13065_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 2 | ||||||||||||
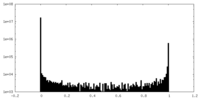
| 投影像・断面図 |
| ||||||||||||
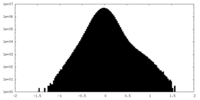
| 密度ヒストグラム |
-ハーフマップ: Half map 1
| ファイル | emd_13065_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 1 | ||||||||||||
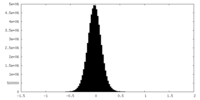
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : ALC1/CHD1L bound to a PARylated nucleosome
| 全体 | 名称: ALC1/CHD1L bound to a PARylated nucleosome |
|---|---|
| 要素 |
|
-超分子 #1: ALC1/CHD1L bound to a PARylated nucleosome
| 超分子 | 名称: ALC1/CHD1L bound to a PARylated nucleosome / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all 詳細: The nucleosome was PARylated by PARP2 and HPF1 before addition of ALC1. |
|---|---|
| 由来(天然) | 生物種: |
| 分子量 | 理論値: 305 KDa |
-分子 #1: Chromodomain-helicase-DNA-binding protein 1-like
| 分子 | 名称: Chromodomain-helicase-DNA-binding protein 1-like / タイプ: protein_or_peptide / ID: 1 / 詳細: C-terminal 6-His tag. / コピー数: 1 / 光学異性体: LEVO / EC番号: DNA helicase |
|---|---|
| 由来(天然) | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 分子量 | 理論値: 98.906766 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MFLLRLHTEG RAEAARVQEQ DLRQWGLTGI HLRSYQLEGV NWLAQRFHCQ NGCILGDEMG LGKTCQTIAL FIYLAGRLND EGPFLILCP LSVLSNWKEE MQRFAPGLSC VTYAGDKEER ACLQQDLKQE SRFHVLLTTY EICLKDASFL KSFPWSVLVV D EAHRLKNQ ...文字列: MFLLRLHTEG RAEAARVQEQ DLRQWGLTGI HLRSYQLEGV NWLAQRFHCQ NGCILGDEMG LGKTCQTIAL FIYLAGRLND EGPFLILCP LSVLSNWKEE MQRFAPGLSC VTYAGDKEER ACLQQDLKQE SRFHVLLTTY EICLKDASFL KSFPWSVLVV D EAHRLKNQ SSLLHKTLSE FSVVFSLLLT GTPIQNSLQE LYSLLSFVEP DLFSKEEVGD FIQRYQDIEK ESESASELHK LL QPFLLRR VKAEVATELP KKTEVVIYHG MSALQKKYYK AILMKDLDAF ENETAKKVKL QNILSQLRKC VDHPYLFDGV EPE PFEVGD HLTEASGKLH LLDKLLAFLY SGGHRVLLFS QMTQMLDILQ DYMDYRGYSY ERVDGSVRGE ERHLAIKNFG QQPI FVFLL STRAGGVGMN LTAADTVIFV DSDFNPQNDL QAAARAHRIG QNKSVKVIRL IGRDTVEEIV YRKAASKLQL TNMII EGGH FTLGAQKPAA DADLQLSEIL KFGLDKLLAS EGSTMDEIDL ESILGETKDG QWVSDALPAA EGGSRDQEEG KNHMYL FEG KDYSKEPSKE DRKSFEQLVN LQKTLLEKAS QEGRSLRNKG SVLIPGLVEG STKRKRVLSP EELEDRQKKR QEAAAKR RR LIEEKKRQKE EAEHKKKMAW WESNNYQSFC LPSEESEPED LENGEESSAE LDYQDPDATS LKYVSGDVTH PQAGAEDA L IVHCVDDSGH WGRGGLFTAL EKRSAEPRKI YELAGKMKDL SLGGVLLFPV DDKESRNKGQ DLLALIVAQH RDRSNVLSG IKMAALEEGL KKIFLAAKKK KASVHLPRIG HATKGFNWYG TERLIRKHLA ARGIPTYIYY FPRSKAHHHH HH UniProtKB: Chromodomain-helicase-DNA-binding protein 1-like |
-分子 #2: Histone H3.2
| 分子 | 名称: Histone H3.2 / タイプ: protein_or_peptide / ID: 2 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種: |
| 分子量 | 理論値: 15.403062 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MARTKQTARK STGGKAPRKQ LATKAARKSA PATGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSSA VMALQEASEA YLVALFEDTN LAAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.2 |
-分子 #3: Histone H4
| 分子 | 名称: Histone H4 / タイプ: protein_or_peptide / ID: 3 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種: |
| 分子量 | 理論値: 11.394426 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-分子 #4: Histone H2A type 1
| 分子 | 名称: Histone H2A type 1 / タイプ: protein_or_peptide / ID: 4 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種: |
| 分子量 | 理論値: 14.109436 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MSGRGKQGGK TRAKAKTRSS RAGLQFPVGR VHRLLRKGNY AERVGAGAPV YLAAVLEYLT AEILELAGNA ARDNKKTRII PRHLQLAVR NDEELNKLLG RVTIAQGGVL PNIQSVLLPK KTESSKSAKS K UniProtKB: Histone H2A type 1 |
-分子 #5: Histone H2B 1.1
| 分子 | 名称: Histone H2B 1.1 / タイプ: protein_or_peptide / ID: 5 / コピー数: 2 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種: |
| 分子量 | 理論値: 13.655948 KDa |
| 組換発現 | 生物種:  |
| 配列 | 文字列: MAKSAPAPKK GSKKAVTKTQ KKDGKKRRKT RKESYAIYVY KVLKQVHPDT GISSKAMSIM NSFVNDVFER IAGEASRLAH YNKRSTITS REIQTAVRLL LPGELAKHAV SEGTKAVTKY TSAK UniProtKB: Histone H2B 1.1 |
-分子 #6: DNA (149-MER) Widom 601 sequence
| 分子 | 名称: DNA (149-MER) Widom 601 sequence / タイプ: dna / ID: 6 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種: synthetic construct (人工物) |
| 分子量 | 理論値: 49.175336 KDa |
| 配列 | 文字列: (DT)(DC)(DT)(DA)(DG)(DG)(DT)(DG)(DA)(DC) (DC)(DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC) (DC)(DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG) (DA)(DG)(DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA) (DA) (DT)(DT)(DG)(DG)(DT) ...文字列: (DT)(DC)(DT)(DA)(DG)(DG)(DT)(DG)(DA)(DC) (DC)(DA)(DT)(DC)(DA)(DG)(DA)(DA)(DT)(DC) (DC)(DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG) (DA)(DG)(DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA) (DA) (DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT) (DA)(DG)(DA)(DC)(DA)(DG)(DC)(DT)(DC)(DT) (DA)(DG) (DC)(DA)(DC)(DC)(DG)(DC)(DT) (DT)(DA)(DA)(DA)(DC)(DG)(DC)(DA)(DC)(DG) (DT)(DA)(DC) (DG)(DC)(DG)(DC)(DT)(DG) (DT)(DC)(DC)(DC)(DC)(DC)(DG)(DC)(DG)(DT) (DT)(DT)(DT)(DA) (DA)(DC)(DC)(DG)(DC) (DC)(DA)(DA)(DG)(DG)(DG)(DG)(DA)(DT)(DT) (DA)(DC)(DT)(DC)(DC) (DC)(DT)(DA)(DG) (DT)(DC)(DT)(DC)(DC)(DA)(DG)(DG)(DC)(DA) (DC)(DG)(DT)(DG)(DT)(DC) (DA)(DG)(DA) (DT)(DA)(DT)(DA)(DT)(DA)(DC)(DA)(DT)(DC) (DG)(DA)(DT)(DA)(DG)(DG)(DC) |
-分子 #7: DNA (149-MER) Widom 601 sequence
| 分子 | 名称: DNA (149-MER) Widom 601 sequence / タイプ: dna / ID: 7 / コピー数: 1 / 分類: DNA |
|---|---|
| 由来(天然) | 生物種: synthetic construct (人工物) |
| 分子量 | 理論値: 49.606586 KDa |
| 配列 | 文字列: (DG)(DC)(DC)(DT)(DA)(DT)(DC)(DG)(DA)(DT) (DG)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DC)(DT) (DG)(DA)(DC)(DA)(DC)(DG)(DT)(DG)(DC) (DC)(DT)(DG)(DG)(DA)(DG)(DA)(DC)(DT)(DA) (DG) (DG)(DG)(DA)(DG)(DT) ...文字列: (DG)(DC)(DC)(DT)(DA)(DT)(DC)(DG)(DA)(DT) (DG)(DT)(DA)(DT)(DA)(DT)(DA)(DT)(DC)(DT) (DG)(DA)(DC)(DA)(DC)(DG)(DT)(DG)(DC) (DC)(DT)(DG)(DG)(DA)(DG)(DA)(DC)(DT)(DA) (DG) (DG)(DG)(DA)(DG)(DT)(DA)(DA)(DT) (DC)(DC)(DC)(DC)(DT)(DT)(DG)(DG)(DC)(DG) (DG)(DT) (DT)(DA)(DA)(DA)(DA)(DC)(DG) (DC)(DG)(DG)(DG)(DG)(DG)(DA)(DC)(DA)(DG) (DC)(DG)(DC) (DG)(DT)(DA)(DC)(DG)(DT) (DG)(DC)(DG)(DT)(DT)(DT)(DA)(DA)(DG)(DC) (DG)(DG)(DT)(DG) (DC)(DT)(DA)(DG)(DA) (DG)(DC)(DT)(DG)(DT)(DC)(DT)(DA)(DC)(DG) (DA)(DC)(DC)(DA)(DA) (DT)(DT)(DG)(DA) (DG)(DC)(DG)(DG)(DC)(DC)(DT)(DC)(DG)(DG) (DC)(DA)(DC)(DC)(DG)(DG) (DG)(DA)(DT) (DT)(DC)(DT)(DG)(DA)(DT)(DG)(DG)(DT)(DC) (DA)(DC)(DC)(DT)(DA)(DG)(DA) |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.3 mg/mL |
|---|---|
| 緩衝液 | pH: 7.5 |
| グリッド | モデル: Quantifoil R2/2 / 材質: COPPER / メッシュ: 200 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY ARRAY / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 60 sec. / 前処理 - 雰囲気: AIR / 前処理 - 気圧: 0.04 kPa / 詳細: Current 20 mA. |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK IV 詳細: 3 uL were applied on grid and immediately blotted for 2.5 s at blot force 0.. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | TFS KRIOS |
|---|---|
| 特殊光学系 | エネルギーフィルター - 名称: GIF Bioquantum / エネルギーフィルター - スリット幅: 20 eV |
| 撮影 | フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) 撮影したグリッド数: 1 / 実像数: 26747 / 平均露光時間: 2.2 sec. / 平均電子線量: 45.0 e/Å2 / 詳細: Total dose was fractionated over 40 movie frames. |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル |
| ||||||
|---|---|---|---|---|---|---|---|
| 精密化 | 空間: REAL / プロトコル: FLEXIBLE FIT / 当てはまり具合の基準: Real-space CC | ||||||
| 得られたモデル |  PDB-7otq: |
 ムービー
ムービー コントローラー
コントローラー
















 Z
Z Y
Y X
X