+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-13009 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Pol II-CSB-CSA-DDB1-UVSSA-ADPBeF3 (Structure2) | |||||||||
 Map data Map data | The main map. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcription / DNA repair | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of double-strand break repair via nonhomologous end joining / nucleotide-excision repair complex / regulation of transcription-coupled nucleotide-excision repair / : / response to auditory stimulus / regulation of transcription elongation by RNA polymerase II / DNA protection / single strand break repair / B-WICH complex / Formation of RNA Pol II elongation complex ...negative regulation of double-strand break repair via nonhomologous end joining / nucleotide-excision repair complex / regulation of transcription-coupled nucleotide-excision repair / : / response to auditory stimulus / regulation of transcription elongation by RNA polymerase II / DNA protection / single strand break repair / B-WICH complex / Formation of RNA Pol II elongation complex / Formation of the Early Elongation Complex / Transcriptional regulation by small RNAs / RNA Polymerase II Pre-transcription Events / TP53 Regulates Transcription of DNA Repair Genes / FGFR2 alternative splicing / RNA polymerase II transcribes snRNA genes / mRNA Capping / mRNA Splicing - Minor Pathway / Processing of Capped Intron-Containing Pre-mRNA / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Elongation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA Pol II CTD phosphorylation and interaction with CE / Estrogen-dependent gene expression / response to superoxide / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / mRNA Splicing - Major Pathway / double-strand break repair via classical nonhomologous end joining / photoreceptor cell maintenance / positive regulation by virus of viral protein levels in host cell / RNA polymerase binding / positive regulation of DNA-templated transcription, elongation / chromatin-protein adaptor activity / spindle assembly involved in female meiosis / response to UV-B / epigenetic programming in the zygotic pronuclei / ATP-dependent chromatin remodeler activity / positive regulation of transcription by RNA polymerase III / UV-damage excision repair / biological process involved in interaction with symbiont / ATP-dependent DNA damage sensor activity / regulation of mitotic cell cycle phase transition / WD40-repeat domain binding / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / Cul4A-RING E3 ubiquitin ligase complex / organelle membrane / Cul4-RING E3 ubiquitin ligase complex / Cul4B-RING E3 ubiquitin ligase complex / positive regulation of transcription by RNA polymerase I / ubiquitin ligase complex scaffold activity / negative regulation of reproductive process / negative regulation of developmental process / RNA Polymerase I Transcription Initiation / protein tyrosine kinase activator activity / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / pyrimidine dimer repair / viral release from host cell / positive regulation of translational initiation / cullin family protein binding / positive regulation of transcription initiation by RNA polymerase II / response to X-ray / ectopic germ cell programmed cell death / ATP-dependent activity, acting on DNA / RNA polymerase I complex / RNA polymerase III complex / positive regulation of viral genome replication / transcription elongation by RNA polymerase I / response to UV / RNA polymerase II, core complex / protein autoubiquitination / tRNA transcription by RNA polymerase III / positive regulation of double-strand break repair via homologous recombination / ubiquitin-like ligase-substrate adaptor activity / transcription by RNA polymerase I / site of DNA damage / JNK cascade / protein localization to chromatin / transcription-coupled nucleotide-excision repair / translation initiation factor binding / proteasomal protein catabolic process / neurogenesis / positive regulation of gluconeogenesis / DNA damage checkpoint signaling / regulation of DNA-templated transcription elongation / positive regulation of RNA splicing / positive regulation of DNA repair / transcription elongation factor complex / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / helicase activity / response to gamma radiation / nucleotide-excision repair / transcription initiation at RNA polymerase II promoter / promoter-specific chromatin binding / sperm end piece / P-body / Recognition of DNA damage by PCNA-containing replication complex Similarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Kokic G / Cramer P | |||||||||
| Funding support |  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Structural basis of human transcription-DNA repair coupling. Authors: Goran Kokic / Felix R Wagner / Aleksandar Chernev / Henning Urlaub / Patrick Cramer /  Abstract: Transcription-coupled DNA repair removes bulky DNA lesions from the genome and protects cells against ultraviolet (UV) irradiation. Transcription-coupled DNA repair begins when RNA polymerase II ...Transcription-coupled DNA repair removes bulky DNA lesions from the genome and protects cells against ultraviolet (UV) irradiation. Transcription-coupled DNA repair begins when RNA polymerase II (Pol II) stalls at a DNA lesion and recruits the Cockayne syndrome protein CSB, the E3 ubiquitin ligase, CRL4 and UV-stimulated scaffold protein A (UVSSA). Here we provide five high-resolution structures of Pol II transcription complexes containing human transcription-coupled DNA repair factors and the elongation factors PAF1 complex (PAF) and SPT6. Together with biochemical and published data, the structures provide a model for transcription-repair coupling. Stalling of Pol II at a DNA lesion triggers replacement of the elongation factor DSIF by CSB, which binds to PAF and moves upstream DNA to SPT6. The resulting elongation complex, EC, uses the CSA-stimulated translocase activity of CSB to pull on upstream DNA and push Pol II forward. If the lesion cannot be bypassed, CRL4 spans over the Pol II clamp and ubiquitylates the RPB1 residue K1268, enabling recruitment of TFIIH to UVSSA and DNA repair. Conformational changes in CRL4 lead to ubiquitylation of CSB and to release of transcription-coupled DNA repair factors before transcription may continue over repaired DNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13009.map.gz emd_13009.map.gz | 95.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13009-v30.xml emd-13009-v30.xml emd-13009.xml emd-13009.xml | 60.5 KB 60.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13009_fsc.xml emd_13009_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_13009.png emd_13009.png | 150.5 KB | ||
| Masks |  emd_13009_msk_1.map emd_13009_msk_1.map emd_13009_msk_2.map emd_13009_msk_2.map emd_13009_msk_3.map emd_13009_msk_3.map emd_13009_msk_4.map emd_13009_msk_4.map | 244.1 MB 244.1 MB 244.1 MB 244.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13009.cif.gz emd-13009.cif.gz | 13.5 KB | ||
| Others |  emd_13009_additional_1.map.gz emd_13009_additional_1.map.gz emd_13009_additional_2.map.gz emd_13009_additional_2.map.gz emd_13009_additional_3.map.gz emd_13009_additional_3.map.gz emd_13009_additional_4.map.gz emd_13009_additional_4.map.gz emd_13009_additional_5.map.gz emd_13009_additional_5.map.gz emd_13009_additional_6.map.gz emd_13009_additional_6.map.gz emd_13009_half_map_1.map.gz emd_13009_half_map_1.map.gz emd_13009_half_map_2.map.gz emd_13009_half_map_2.map.gz | 195.8 MB 195.8 MB 194 MB 193.8 MB 194.6 MB 193.8 MB 221 MB 220.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13009 http://ftp.pdbj.org/pub/emdb/structures/EMD-13009 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13009 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13009 | HTTPS FTP |
-Related structure data
| Related structure data |  7oobMC  7oo3C  7oopC  7opcC  7opdC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13009.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13009.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | The main map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
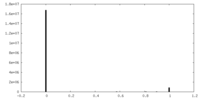
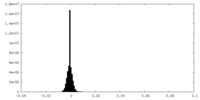
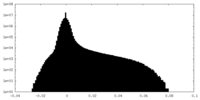
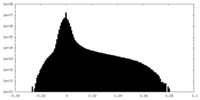
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
+Mask #1
+Mask #2
+Mask #3
+Mask #4
+Additional map: Half map corresponding to the map focused refined on Pol II.
+Additional map: Half map corresponding to the map focused refined on Pol II.
+Additional map: Half map corresponding to the map focused refined on CSA-DDB1.
+Additional map: Half map corresponding to the map focused refined on CSA-DDB1.
+Additional map: Half map corresponding to the map focused refined on CSB.
+Additional map: Half map corresponding to the map focused refined on CSB.
+Half map: Half map corresponding to the main map.
+Half map: Half map corresponding to the main map.
- Sample components
Sample components
+Entire : Complex between the transcribing RNA polymerase II and TCR factor...
+Supramolecule #1: Complex between the transcribing RNA polymerase II and TCR factor...
+Supramolecule #2: DNA-directed RNA polymerase with RNA_pol_L_2 and RNA polymerase
+Supramolecule #3: DNA excision repair protein ERCC-6 and ERCC-8, DNA damage-binding...
+Supramolecule #4: NTS, RNA, TS
+Macromolecule #1: DNA-directed RNA polymerase II subunit RPB1
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase II subunit RPB3
+Macromolecule #4: RPOL4c domain-containing protein
+Macromolecule #5: DNA-directed RNA polymerase II subunit E
+Macromolecule #6: DNA-directed RNA polymerase II subunit F
+Macromolecule #7: DNA-directed RNA polymerase II subunit RPB7
+Macromolecule #8: DNA-directed RNA polymerases I, II, and III subunit RPABC3
+Macromolecule #9: DNA-directed RNA polymerase II subunit RPB9
+Macromolecule #10: DNA-directed RNA polymerases I, II, and III subunit RPABC5
+Macromolecule #11: RNA_pol_L_2 domain-containing protein
+Macromolecule #12: RNA polymerase II subunit K
+Macromolecule #13: DNA excision repair protein ERCC-6
+Macromolecule #14: DNA damage-binding protein 1
+Macromolecule #18: DNA excision repair protein ERCC-8
+Macromolecule #15: NTS
+Macromolecule #16: TS
+Macromolecule #17: RNA
+Macromolecule #19: ZINC ION
+Macromolecule #20: MAGNESIUM ION
+Macromolecule #21: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #22: BERYLLIUM TRIFLUORIDE ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/1 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 10940 / Average electron dose: 41.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)

















































































































 Trichoplusia ni (cabbage looper)
Trichoplusia ni (cabbage looper)


