+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11854 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the human THO - UAP56 complex (Map C) | |||||||||
 Map data Map data | Map C sharpened (Map sharpening B-factor -160 A%u030A2) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.7 Å | |||||||||
 Authors Authors | Puehringer C / Hohmann U / Plaschka C | |||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Structure of the human core transcription-export complex reveals a hub for multivalent interactions. Authors: Thomas Pühringer / Ulrich Hohmann / Laura Fin / Belén Pacheco-Fiallos / Ulla Schellhaas / Julius Brennecke / Clemens Plaschka /  Abstract: The export of mRNA from nucleus to cytoplasm requires the conserved and essential transcription and export (TREX) complex (THO-UAP56/DDX39B-ALYREF). TREX selectively binds mRNA maturation marks and ...The export of mRNA from nucleus to cytoplasm requires the conserved and essential transcription and export (TREX) complex (THO-UAP56/DDX39B-ALYREF). TREX selectively binds mRNA maturation marks and licenses mRNA for nuclear export by loading the export factor NXF1-NXT1. How TREX integrates these marks and achieves high selectivity for mature mRNA is poorly understood. Here, we report the cryo-electron microscopy structure of the human THO-UAP56/DDX39B complex at 3.3 Å resolution. The seven-subunit THO-UAP56/DDX39B complex multimerizes into a 28-subunit tetrameric assembly, suggesting that selective recognition of mature mRNA is facilitated by the simultaneous sensing of multiple, spatially distant mRNA regions and maturation marks. Two UAP56/DDX39B RNA helicases are juxtaposed at each end of the tetramer, which would allow one bivalent ALYREF protein to bridge adjacent helicases and regulate the TREX-mRNA interaction. Our structural and biochemical results suggest a conserved model for TREX complex function that depends on multivalent interactions between proteins and mRNA. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11854.map.gz emd_11854.map.gz | 303.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11854-v30.xml emd-11854-v30.xml emd-11854.xml emd-11854.xml | 12.3 KB 12.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11854.png emd_11854.png | 13.7 KB | ||
| Others |  emd_11854_additional_1.map.gz emd_11854_additional_1.map.gz | 300.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11854 http://ftp.pdbj.org/pub/emdb/structures/EMD-11854 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11854 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11854 | HTTPS FTP |
-Validation report
| Summary document |  emd_11854_validation.pdf.gz emd_11854_validation.pdf.gz | 253 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11854_full_validation.pdf.gz emd_11854_full_validation.pdf.gz | 252.1 KB | Display | |
| Data in XML |  emd_11854_validation.xml.gz emd_11854_validation.xml.gz | 7.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11854 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11854 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11854 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11854 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11854.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11854.map.gz / Format: CCP4 / Size: 325 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map C sharpened (Map sharpening B-factor -160 A%u030A2) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
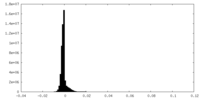
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Map C unsharpened
| File | emd_11854_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Map C unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
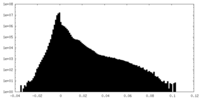
| Density Histograms |
- Sample components
Sample components
-Entire : THO - UAP56
| Entire | Name: THO - UAP56 |
|---|---|
| Components |
|
-Supramolecule #1: THO - UAP56
| Supramolecule | Name: THO - UAP56 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#14 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) / Recombinant strain: Hi5 Trichoplusia ni (cabbage looper) / Recombinant strain: Hi5 |
| Molecular weight | Theoretical: 1.836 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 2 / Number real images: 26303 / Average exposure time: 3.8 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)