+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10103 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of murine norovirus (MNV-1) | ||||||||||||
 Map data Map data | None | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Norovirus / Virus / Capsid / VP1 | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationantigenic variation / symbiont-mediated perturbation of host defense response / virion component / host cell cytoplasm Similarity search - Function | ||||||||||||
| Biological species |   Murine norovirus 1 Murine norovirus 1 | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | ||||||||||||
 Authors Authors | Snowden JS / Hurdiss DL | ||||||||||||
| Funding support |  United Kingdom, 3 items United Kingdom, 3 items
| ||||||||||||
 Citation Citation |  Journal: PLoS Biol / Year: 2020 Journal: PLoS Biol / Year: 2020Title: Dynamics in the murine norovirus capsid revealed by high-resolution cryo-EM. Authors: Joseph S Snowden / Daniel L Hurdiss / Oluwapelumi O Adeyemi / Neil A Ranson / Morgan R Herod / Nicola J Stonehouse /  Abstract: Icosahedral viral capsids must undergo conformational rearrangements to coordinate essential processes during the viral life cycle. Capturing such conformational flexibility has been technically ...Icosahedral viral capsids must undergo conformational rearrangements to coordinate essential processes during the viral life cycle. Capturing such conformational flexibility has been technically challenging yet could be key for developing rational therapeutic agents to combat infections. Noroviruses are nonenveloped, icosahedral viruses of global importance to human health. They are a common cause of acute gastroenteritis, yet no vaccines or specific antiviral agents are available. Here, we use genetics and cryo-electron microscopy (cryo-EM) to study the high-resolution solution structures of murine norovirus as a model for human viruses. By comparing our 3 structures (at 2.9- to 3.1-Å resolution), we show that whilst there is little change to the shell domain of the capsid, the radiating protruding domains are flexible, adopting distinct states both independently and synchronously. In doing so, the capsids sample a range of conformational space, with implications for maintaining virion stability and infectivity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10103.map.gz emd_10103.map.gz | 261.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10103-v30.xml emd-10103-v30.xml emd-10103.xml emd-10103.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10103_fsc.xml emd_10103_fsc.xml | 17.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_10103.png emd_10103.png | 198.3 KB | ||
| Masks |  emd_10103_msk_1.map emd_10103_msk_1.map | 476.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10103.cif.gz emd-10103.cif.gz | 6.3 KB | ||
| Others |  emd_10103_half_map_1.map.gz emd_10103_half_map_1.map.gz emd_10103_half_map_2.map.gz emd_10103_half_map_2.map.gz | 376.7 MB 376.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10103 http://ftp.pdbj.org/pub/emdb/structures/EMD-10103 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10103 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10103 | HTTPS FTP |
-Related structure data
| Related structure data |  6s6lMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10103.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10103.map.gz / Format: CCP4 / Size: 476.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.065 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10103_msk_1.map emd_10103_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
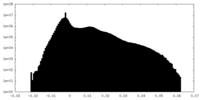
| Density Histograms |
-Half map: #1
| File | emd_10103_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_10103_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Murine norovirus 1
| Entire | Name:   Murine norovirus 1 Murine norovirus 1 |
|---|---|
| Components |
|
-Supramolecule #1: Murine norovirus 1
| Supramolecule | Name: Murine norovirus 1 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / Details: RAW264.7 cells / NCBI-ID: 223997 / Sci species name: Murine norovirus 1 / Sci species strain: CW1P3 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  |
| Virus shell | Shell ID: 1 / T number (triangulation number): 3 |
-Macromolecule #1: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Murine norovirus 1 Murine norovirus 1 |
| Molecular weight | Theoretical: 58.700598 KDa |
| Sequence | String: MRMSDGAAPK ANGSEASGQD LVPAAVEQAV PIQPVAGAAL AAPAAGQINQ IDPWIFQNFV QCPLGEFSIS PRNTPGEILF DLALGPGLN PYLAHLSAMY TGWVGNMEVQ LVLAGNAFTA GKVVVALVPP YFPKGSLTTA QITCFPHVMC DVRTLEPIQL P LLDVRRVL ...String: MRMSDGAAPK ANGSEASGQD LVPAAVEQAV PIQPVAGAAL AAPAAGQINQ IDPWIFQNFV QCPLGEFSIS PRNTPGEILF DLALGPGLN PYLAHLSAMY TGWVGNMEVQ LVLAGNAFTA GKVVVALVPP YFPKGSLTTA QITCFPHVMC DVRTLEPIQL P LLDVRRVL WHATQDQEES MRLVCMLYTP LRTNSPGDES FVVSGRLLSK PAADFNFVYL TPPIERTIYR MVDLPVIQPR LC THARWPA PVYGLLVDPS LPSNPQWQNG RVHVDGTLLG TTPISGSWVS CFAAEAAYEF QSGTGEVATF TLIEQDGSAY VPG DRAAPL GYPDFSGQLE IEVQTETTKT GDKLKVTTFE MILGPTTNAD QAPYQGRVFA SVTAAASLDL VDGRVRAVPR SIYG FQDTI PEYNDGLLVP LAPPIGPFLP GEVLLRFRTY MRQIDTADAA AEAIDCALPQ EFVSWFASNA FTVQSEALLL RYRNT LTGQ LLFECKLYNE GYIALSYSGS GPLTFPTDGI FEVVSWVPRL YQLASVGSLA TGRMLKQ UniProtKB: Capsid protein VP1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Component:
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Material: COPPER / Mesh: 400 | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 80 % / Chamber temperature: 281 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 59.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)