[English] 日本語
 Yorodumi
Yorodumi- EMDB-0345: Metabotropic Glutamate Receptor 5 bound to L-quisqualate and Nb43 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0345 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Metabotropic Glutamate Receptor 5 bound to L-quisqualate and Nb43 | |||||||||
 Map data Map data | Metabotropic Glutamate Receptor 5 bound to L-quisqualate and Nb43, primary map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cell Surface Receptor / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationA2A adenosine receptor binding / sensory perception of hot stimulus / negative regulation of dendritic spine morphogenesis / : / G protein-coupled receptor activity involved in regulation of postsynaptic membrane potential / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / operant conditioning / protein localization to nuclear inner membrane / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / positive regulation of cellular response to hypoxia ...A2A adenosine receptor binding / sensory perception of hot stimulus / negative regulation of dendritic spine morphogenesis / : / G protein-coupled receptor activity involved in regulation of postsynaptic membrane potential / adenylate cyclase inhibiting G protein-coupled glutamate receptor activity / operant conditioning / protein localization to nuclear inner membrane / phospholipase C-activating G protein-coupled glutamate receptor signaling pathway / positive regulation of cellular response to hypoxia / positive regulation of long-term neuronal synaptic plasticity / positive regulation of sensory perception of pain / negative regulation of excitatory postsynaptic potential / positive regulation of dopamine secretion / desensitization of G protein-coupled receptor signaling pathway / G protein-coupled glutamate receptor signaling pathway / positive regulation of neural precursor cell proliferation / Class C/3 (Metabotropic glutamate/pheromone receptors) / nuclear inner membrane / glutamate receptor activity / Neurexins and neuroligins / astrocyte projection / response to corticosterone / response to morphine / temperature homeostasis / protein tyrosine kinase activator activity / conditioned place preference / regulation of synaptic transmission, glutamatergic / regulation of long-term synaptic depression / positive regulation of calcium-mediated signaling / protein tyrosine kinase binding / dendritic shaft / response to amphetamine / locomotory behavior / synapse organization / postsynaptic density membrane / G protein-coupled receptor activity / Schaffer collateral - CA1 synapse / cognition / cellular response to amyloid-beta / positive regulation of cytosolic calcium ion concentration / dendritic spine / G alpha (q) signalling events / chemical synaptic transmission / response to ethanol / learning or memory / positive regulation of MAPK cascade / response to antibiotic / neuronal cell body / regulation of DNA-templated transcription / dendrite / glutamatergic synapse / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.0 Å | |||||||||
 Authors Authors | Koehl A / Hu H | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Structural insights into the activation of metabotropic glutamate receptors. Authors: Antoine Koehl / Hongli Hu / Dan Feng / Bingfa Sun / Yan Zhang / Michael J Robertson / Matthew Chu / Tong Sun Kobilka / Toon Laeremans / Jan Steyaert / Jeffrey Tarrasch / Somnath Dutta / ...Authors: Antoine Koehl / Hongli Hu / Dan Feng / Bingfa Sun / Yan Zhang / Michael J Robertson / Matthew Chu / Tong Sun Kobilka / Toon Laeremans / Jan Steyaert / Jeffrey Tarrasch / Somnath Dutta / Rasmus Fonseca / William I Weis / Jesper M Mathiesen / Georgios Skiniotis / Brian K Kobilka /     Abstract: Metabotropic glutamate receptors are family C G-protein-coupled receptors. They form obligate dimers and possess extracellular ligand-binding Venus flytrap domains, which are linked by cysteine-rich ...Metabotropic glutamate receptors are family C G-protein-coupled receptors. They form obligate dimers and possess extracellular ligand-binding Venus flytrap domains, which are linked by cysteine-rich domains to their 7-transmembrane domains. Spectroscopic studies show that signalling is a dynamic process, in which large-scale conformational changes underlie the transmission of signals from the extracellular Venus flytraps to the G protein-coupling domains-the 7-transmembrane domains-in the membrane. Here, using a combination of X-ray crystallography, cryo-electron microscopy and signalling studies, we present a structural framework for the activation mechanism of metabotropic glutamate receptor subtype 5. Our results show that agonist binding at the Venus flytraps leads to a compaction of the intersubunit dimer interface, thereby bringing the cysteine-rich domains into close proximity. Interactions between the cysteine-rich domains and the second extracellular loops of the receptor enable the rigid-body repositioning of the 7-transmembrane domains, which come into contact with each other to initiate signalling. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0345.map.gz emd_0345.map.gz | 59.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0345-v30.xml emd-0345-v30.xml emd-0345.xml emd-0345.xml | 16.9 KB 16.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0345.png emd_0345.png | 90.3 KB | ||
| Filedesc metadata |  emd-0345.cif.gz emd-0345.cif.gz | 6.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0345 http://ftp.pdbj.org/pub/emdb/structures/EMD-0345 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0345 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0345 | HTTPS FTP |
-Related structure data
| Related structure data |  6n51MC  0346C  0347C  6n4xC  6n4yC  6n50C  6n52C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0345.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0345.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Metabotropic Glutamate Receptor 5 bound to L-quisqualate and Nb43, primary map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Complex between metabotropic glutamate receptor 5 bound to L-quis...
| Entire | Name: Complex between metabotropic glutamate receptor 5 bound to L-quisqualate and Nb43 |
|---|---|
| Components |
|
-Supramolecule #1: Complex between metabotropic glutamate receptor 5 bound to L-quis...
| Supramolecule | Name: Complex between metabotropic glutamate receptor 5 bound to L-quisqualate and Nb43 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 220 KDa |
-Macromolecule #1: Metabotropic glutamate receptor 5
| Macromolecule | Name: Metabotropic glutamate receptor 5 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 90.484805 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: SERRVVAHMP GDIIIGALFS VHHQPTVDKV HERKCGAVRE QYGIQRVEAM LHTLERINSD PTLLPNITLG CEIRDSCWHS AVALEQSIE FIRDSLISSE EEEGLVRCVD GSSSSFRSKK PIVGVIGPGS SSVAIQVQNL LQLFNIPQIA YSATSMDLSD K TLFKYFMR ...String: SERRVVAHMP GDIIIGALFS VHHQPTVDKV HERKCGAVRE QYGIQRVEAM LHTLERINSD PTLLPNITLG CEIRDSCWHS AVALEQSIE FIRDSLISSE EEEGLVRCVD GSSSSFRSKK PIVGVIGPGS SSVAIQVQNL LQLFNIPQIA YSATSMDLSD K TLFKYFMR VVPSDAQQAR AMVDIVKRYN WTYVSAVHTE GNYGESGMEA FKDMSAKEGI CIAHSYKIYS NAGEQSFDKL LK KLTSHLP KARVVACFCE GMTVRGLLMA MRRLGLAGEF LLLGSDGWAD RYDVTDGYQR EAVGGITIKL QSPDVKWFDD YYL KLRPET NHRNPWFQEF WQHRFQCRLE GFPQENSKYN KTCNSSLTLK THHVQDSKMG FVINAIYSMA YGLHNMQMSL CPGY AGLCD AMKPIDGRKL LESLMKTNFT GVSGDTILFD ENGDSPGRYE IMNFKEMGKD YFDYINVGSW DNGELKMDDD EVWSK KSNI IRSVCSEPCE KGQIKVIRKG EVSCCWTCTP CKENEYVFDE YTCKACQLGS WPTDDLTGCD LIPVQYLRWG DPEPIA AVV FACLGLLATL FVTVVFIIYR DTPVVKSSSR ELCYIILAGI CLGYLCTFCL IAKPKQIYCY LQRIGIGLSP AMSYSAL VT KTNRIARILA GSKKKICTKK PRFMSACAQL VIAFILICIQ LGIIVALFIM EPPDIMHDYP SIREVYLICN TTNLGVVT P LGYNGLLILS CTFYAFKTRN VPANFNEAKY IAFTMYTTCI IWLAFVPIYF GSNYKIITMC FSVSLSATVA LGCMFVPKV YIIL UniProtKB: Metabotropic glutamate receptor 5 |
-Macromolecule #2: Nanobody 43
| Macromolecule | Name: Nanobody 43 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.354672 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QVQLVESGGG LVQAGGSLRL SCAASGRTFT SYAMGWFRQA PGKERESVAA ISSSGGSTHY ADSVKGRFTI SRDNSKNTVY LQMNSLKPE DTAVYYCAAA MYGSRWPDWE YDYWGQGTQV TVSS |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 4 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: (S)-2-AMINO-3-(3,5-DIOXO-[1,2,4]OXADIAZOLIDIN-2-YL)-PROPIONIC ACID
| Macromolecule | Name: (S)-2-AMINO-3-(3,5-DIOXO-[1,2,4]OXADIAZOLIDIN-2-YL)-PROPIONIC ACID type: ligand / ID: 4 / Number of copies: 2 / Formula: QUS |
|---|---|
| Molecular weight | Theoretical: 189.126 Da |
| Chemical component information | 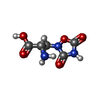 ChemComp-QUS: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 12 mg/mL | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 80 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 390.0 kPa | |||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK IV Details: additional 0.1%OG was added to the sample right before applying sample to grid; 3.5ul sample was applied; blot for 1 second before plunging;. | |||||||||||||||
| Details | Sample was mono disperse as assayed by size exclusion chromatography |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 8.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 47169 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: Ab initio map was generated by VIPER. |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 4.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 2.1) / Number images used: 73472 |
| Initial angle assignment | Type: RANDOM ASSIGNMENT / Software - Name: cryoSPARC |
| Final angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION (ver. 2.1) |
-Atomic model buiding 1
| Refinement | Protocol: FLEXIBLE FIT / Target criteria: Geometry |
|---|---|
| Output model |  PDB-6n51: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















