+Search query
-Structure paper
| Title | Insights into the modulation of bacterial NADase activity by phage proteins. |
|---|---|
| Journal, issue, pages | Nat Commun, Vol. 15, Issue 1, Page 2692, Year 2024 |
| Publish date | Mar 27, 2024 |
 Authors Authors | Hang Yin / Xuzichao Li / Xiaoshen Wang / Chendi Zhang / Jiaqi Gao / Guimei Yu / Qiuqiu He / Jie Yang / Xiang Liu / Yong Wei / Zhuang Li / Heng Zhang /  |
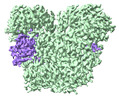
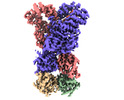
| PubMed Abstract | The Silent Information Regulator 2 (SIR2) protein is widely implicated in antiviral response by depleting the cellular metabolite NAD. The defense-associated sirtuin 2 (DSR2) effector, a SIR2 domain- ...The Silent Information Regulator 2 (SIR2) protein is widely implicated in antiviral response by depleting the cellular metabolite NAD. The defense-associated sirtuin 2 (DSR2) effector, a SIR2 domain-containing protein, protects bacteria from phage infection by depleting NAD, while an anti-DSR2 protein (DSR anti-defense 1, DSAD1) is employed by some phages to evade this host defense. The NADase activity of DSR2 is unleashed by recognizing the phage tail tube protein (TTP). However, the activation and inhibition mechanisms of DSR2 are unclear. Here, we determine the cryo-EM structures of DSR2 in multiple states. DSR2 is arranged as a dimer of dimers, which is facilitated by the tetramerization of SIR2 domains. Moreover, the DSR2 assembly is essential for activating the NADase function. The activator TTP binding would trigger the opening of the catalytic pocket and the decoupling of the N-terminal SIR2 domain from the C-terminal domain (CTD) of DSR2. Importantly, we further show that the activation mechanism is conserved among other SIR2-dependent anti-phage systems. Interestingly, the inhibitor DSAD1 mimics TTP to trap DSR2, thus occupying the TTP-binding pocket and inhibiting the NADase function. Together, our results provide molecular insights into the regulatory mechanism of SIR2-dependent NAD depletion in antiviral immunity. |
 External links External links |  Nat Commun / Nat Commun /  PubMed:38538592 / PubMed:38538592 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 2.9 - 3.9 Å |
| Structure data | EMDB-36980, PDB-8k98: EMDB-36982, PDB-8k9a: EMDB-37272, PDB-8w56: EMDB-37603, PDB-8wkn: EMDB-38421, PDB-8xkn: |
| Source |
|
 Keywords Keywords | ANTIVIRAL PROTEIN / cryo-EM structure of a protein |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers