-検索条件
-検索結果
検索 (著者・登録者: valle & m)の結果200件中、1から50件目までを表示しています

EMDB-17349: 
Human N-deacetylase/N-sulfotransferase 1 homodimer

EMDB-41153: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf

EMDB-41154: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf

PDB-8tcf: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf

PDB-8tcg: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf

EMDB-15036: 
Cryo-EM structure of "CT-CT dimer" of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz6: 
Cryo-EM structure of "CT-CT dimer" of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15028: 
Cryo-EM structure of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15029: 
Cryo-EM structure of "CT oxa" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15030: 
Cryo-EM structure of "CT empty" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15031: 
Cryo-EM structure of "CT react" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15032: 
Cryo-EM structure of "CT pyr" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15033: 
Cryo-EM structure of "BC react" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15034: 
Cryo-EM structure of "BC closed" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15035: 
Cryo-EM structure of "BC open" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

EMDB-15037: 
Cryo-EM structure of Lactococcus lactis pyruvate carboxylase with acetyl-CoA and cyclic di-AMP

PDB-7zyy: 
Cryo-EM structure of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zyz: 
Cryo-EM structure of "CT oxa" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz0: 
Cryo-EM structure of "CT empty" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz1: 
Cryo-EM structure of "CT react" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz2: 
Cryo-EM structure of "CT pyr" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz3: 
Cryo-EM structure of "BC react" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz4: 
Cryo-EM structure of "BC closed" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz5: 
Cryo-EM structure of "BC open" conformation of Lactococcus lactis pyruvate carboxylase with acetyl-CoA

PDB-7zz8: 
Cryo-EM structure of Lactococcus lactis pyruvate carboxylase with acetyl-CoA and cyclic di-AMP

EMDB-26874: 
Band 3-Glycophorin A complex, outward facing

EMDB-26886: 
Erythrocyte ankyrin-1 complex class 2 local refinement of AQP1 (C4 symmetry applied)

EMDB-26916: 
Local refinement of RhAG-RhCE-ANK1(AR1-5), from consensus refinement of all classes

EMDB-26917: 
Protein 4.2 (local refinement from consensus reconstruction of ankyrin complex classes)
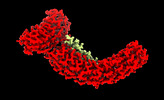
EMDB-26918: 
Ankyrin-1 (N-terminal region of membrane binding domain, local refinement from consensus reconstruction; bound to N-terminal peptide from band 3)

EMDB-26919: 
Cytoplasmic domains of Band 3-I (local refinement from consensus reconstruction of ankyrin complexes)

EMDB-26940: 
Band 3-I-TM local refinement from erythrocyte ankyrin-1 complex consensus reconstruction
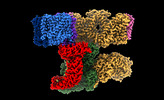
EMDB-26943: 
Consensus refinement of human erythrocyte ankyrin-1 complex (Composite map)

EMDB-26944: 
Local refinement of ankyrin-1 (N-terminal half), class 1 of erythrocyte ankyrin-1 complex

EMDB-26948: 
Local refinement of protein 4.2, class 1 of erythrocyte ankyrin-1 complex

EMDB-26949: 
Local refinement of RhAG/CE trimer, class 1 of erythrocyte ankyrin-1 complex

EMDB-26950: 
Local refinement of Band 3-I cytoplasmic domains, class 1 of erythrocyte ankyrin-1 complex

EMDB-26951: 
Local refinement of Band 3-II cytoplasmic domains, class 1 of erythrocyte ankyrin-1 complex
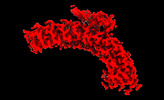
EMDB-26952: 
Local refinement of ankyrin-1 (C-terminal half), class 1 of erythrocyte ankyrin-1 complex

EMDB-26953: 
Local refinement of Band 3-III cytoplasmic domains, class 1 of erythrocyte ankyrin-1 complex

EMDB-26954: 
Local refinement of Band 3-II transmembrane domains, class 1 of erythrocyte ankyrin-1 complex

EMDB-26955: 
Local refinement of Band 3-I transmembrane domains, class 1 of erythrocyte ankyrin-1 complex
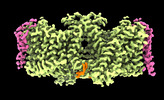
EMDB-26956: 
Local refinement of Band 3-III transmembrane domains, class 1 of erythrocyte ankyrin-1 complex

EMDB-26958: 
Local refinement of Rh trimer, glycophorin B and Band3-III transmembrane region, class 1a of erythrocyte ankyrin-1 complex

EMDB-26960: 
Composite reconstruction of Class 1 of the erythrocyte ankyrin-1 complex

EMDB-26965: 
Sub-tomogram averaging of erythrocyte ankyrin-1 complex
EMDB-26972: 
Local refinement of Anykyrin-1 (N-terminal half of membrane binding domain) in Class 2 of erythrocyte ankyrin-1 complex

EMDB-26973: 
Local refinement of protein 4.2 in Class 2 of erythrocyte ankyrin-1 complex

EMDB-26974: 
Local refinement of RhAG/CE trimer in class 2 of erythrocyte ankyrin-1 complex

EMDB-26975: 
Local refinement of cytoplasmic domains of band3-I in class 2 of erythrocyte ankyrin-1 complex
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
