-Search query
-Search result
Showing 1 - 50 of 187 items for (author: torres & ci)

EMDB-29330: 
N332-GT5 SOSIP in complex with base polyclonal Fabs isolated at day 42 from protein immunized wild type mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB

EMDB-29333: 
N332-GT5 SOSIP in complex with base polyclonal Fabs isolated at day 42 from protein immunized BG18HCgl knock-in mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB

EMDB-29334: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 16 from mRNA immunized wild type mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB

EMDB-29335: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 42 from mRNA immunized BG18HCgl knock-in mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB

EMDB-43191: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 14 from N332-GT2 nanoparticle-immunized BG18HCgl knock-in mice
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-43192: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 15 from N332-GT5 nanoparticle-immunized BG18HCgl knock-in mice
Method: single particle / : Ozorowski G, Torres JL, Ward AB
EMDB-28937: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 42 from protein immunized wild type mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB
EMDB-28938: 
N332-GT5 SOSIP in complex with V1V3 and base polyclonal Fabs isolated at day 16 from protein immunized BG18HCgl knock-in mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB
EMDB-28939: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 42 from mRNA immunized wild type mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB
EMDB-28940: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 16 from mRNA immunized BG18HCgl knock-in mice
Method: single particle / : Torres JL, Jackson AM, Ozorowski G, Ward AB

EMDB-28941: 
HIV Env BG505_MD39_B11 SOSIP boosting trimer in complex with B11_d77.7 mouse Fab and RM20A3 Fab
Method: single particle / : Torres JL, Ozorowski G, Ward AB

EMDB-28942: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized BG18HCgl knock-in mice
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-28945: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized wild type mice, and NHP monoclonal Fab RM20A3
Method: single particle / : Ozorowski G, Ward AB

EMDB-43190: 
HIV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab
Method: single particle / : Ozorowski G, Torres JL, Ward AB

PDB-8f92: 
HIV Env BG505_MD39_B11 SOSIP boosting trimer in complex with B11_d77.7 mouse Fab and RM20A3 Fab
Method: single particle / : Torres JL, Ozorowski G, Ward AB

PDB-8f9g: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized BG18HCgl knock-in mice
Method: single particle / : Ozorowski G, Torres JL, Ward AB

PDB-8f9m: 
HIV Env germline targeting BG505_MD64_N332-GT5 SOSIP in complex with V3-glycan polyclonal Fab isolated from immunized wild type mice, and NHP monoclonal Fab RM20A3
Method: single particle / : Ozorowski G, Ward AB

PDB-8vfv: 
HIV Env BG505_MD39_B16 SOSIP boosting trimer in complex with B16_d77.5 mouse Fab and RM20A3 Fab
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vye: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyf: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyg: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-27781: 
Cryo-EM structure of 227 Fab in complex with (NPNA)8 peptide
Method: single particle / : Martin GM, Ward AB

EMDB-27784: 
Cryo-EM structure of 239 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27785: 
Cryo-EM structure of 311 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27786: 
Cryo-EM structure of 334 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27787: 
Cryo-EM structure of 337 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27788: 
Cryo-EM structure of 356 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

EMDB-27789: 
Cryo-EM structure of 364 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dyt: 
Cryo-EM structure of 227 Fab in complex with (NPNA)8 peptide
Method: single particle / : Martin GM, Ward AB

PDB-8dyw: 
Cryo-EM structure of 239 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dyx: 
Cryo-EM structure of 311 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dyy: 
Cryo-EM structure of 334 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dz3: 
Cryo-EM structure of 337 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dz4: 
Cryo-EM structure of 356 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB

PDB-8dz5: 
Cryo-EM structure of 364 Fab in complex with recombinant shortened Plasmodium falciparum circumsporozoite protein (rsCSP)
Method: single particle / : Martin GM, Ward AB
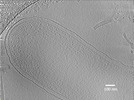
EMDB-27710: 
Intracytoplasmic membranes in Geobacter sulfurreducens
Method: electron tomography / : Howley EH, Williams DR, Torres CI
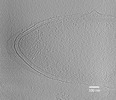
EMDB-27729: 
Intracytoplasmic membranes in a tomogram of Geobacter sulfurreducens
Method: electron tomography / : Howley EH, Williams DR, Torres CI
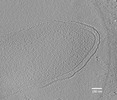
EMDB-27747: 
Electron cryotomogram of Geobacter sulfurreducens cell.
Method: electron tomography / : Howley ET, Williams DR, Torres CI
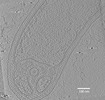
EMDB-27748: 
Geobacter sulfurreducens whole cell tomogram with membrane invagination
Method: electron tomography / : Howley ET, Williams DR, Torres CI

EMDB-15039: 
BcsH-BcsD 'beads-on-a-string' filament, local refine
Method: single particle / : Krasteva PV, Abidi W, Decossas M

EMDB-15040: 
BcsHD cis-filaments: four 'beads-on-a-string'
Method: single particle / : Krasteva PV, Abidi W, Decossas M

EMDB-15041: 
Solution BcsD structure
Method: single particle / : Krasteva PV, Abidi W, Decossas M

PDB-7zzq: 
BcsH-BcsD 'beads-on-a-string' filament, local refine
Method: single particle / : Krasteva PV, Abidi W, Decossas M

PDB-7zzy: 
Solution BcsD structure
Method: single particle / : Krasteva PV, Abidi W, Decossas M

EMDB-27610: 
BG505 MD39 SOSIP in complex with Rh.NJ82 wk13 N611 and base epitope pAbs
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
EMDB-27611: 
BG505 MD39 SOSIP in complex with Rh.NJ95 wk13 base epitope pAb
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
EMDB-27612: 
BG505 MD39 SOSIP in complex with Rh.NK05 wk13 base epitope pAb
Method: single particle / : Torres JL, Lee WH, Ozorowski G, Ward AB
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
