-Search query
-Search result
Showing 1 - 50 of 382 items for (author: smith & ga)

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

PDB-8sx3: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

EMDB-41024: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

EMDB-41025: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

EMDB-41026: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

EMDB-41027: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

EMDB-41034: 
MD64 N332-GT5 SOSIP

EMDB-41035: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

PDB-8t49: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

PDB-8t4a: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

PDB-8t4b: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

PDB-8t4d: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

PDB-8t4k: 
MD64 N332-GT5 SOSIP

PDB-8t4l: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

EMDB-42977: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)

EMDB-43000: 
Cryo-EM structure of SNF2h-nucleosome complex (consensus structure)

EMDB-43001: 
Cryo-EM structure of SNF2h-nucleosome complex (single-bound structure)

EMDB-43002: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex

EMDB-43003: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)

EMDB-43004: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 1)

EMDB-43005: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex (conformation 2)

PDB-8v4y: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 1)

PDB-8v6v: 
Cryo-EM structure of doubly-bound SNF2h-nucleosome complex

PDB-8v7l: 
Cryo-EM structure of singly-bound SNF2h-nucleosome complex with SNF2h at inactive SHL2 (conformation 2)
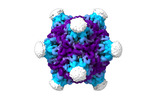
EMDB-19024: 
Structure of the PNMA2 capsid
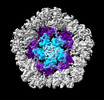
EMDB-19025: 
Structure of the five-fold capsomer of the PNMA2 capsid
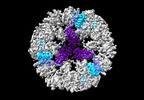
EMDB-19026: 
Structure of the three-fold capsomer of the PNMA2 capsid
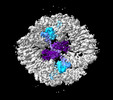
EMDB-19027: 
Structure of the two-fold capsomer of the PNMA2 capsid

EMDB-19194: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #2
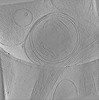

EMDB-19346: 
In situ cryo-electron tomogram of an autophagosome in the projection of an iPSC-derived neuron #1

EMDB-40334: 
Human OCT1 (Apo) in inward-open conformation

EMDB-40335: 
Human OCT1 bound to diltiazem in inward-open conformation

EMDB-40336: 
Human OCT1 bound to fenoterol in inward-open conformation

EMDB-40337: 
Human OCT1 bound to metformin in inward-open conformation

EMDB-40339: 
Human OCT1 bound to thiamine in inward-open conformation

PDB-8sc1: 
Human OCT1 (Apo) in inward-open conformation

PDB-8sc2: 
Human OCT1 bound to diltiazem in inward-open conformation

PDB-8sc3: 
Human OCT1 bound to fenoterol in inward-open conformation

PDB-8sc4: 
Human OCT1 bound to metformin in inward-open conformation

PDB-8sc6: 
Human OCT1 bound to thiamine in inward-open conformation

EMDB-16110: 
Human Urea Transporter UT-A (N-Terminal Domain Model)

EMDB-16111: 
Map of Human Urea Transporter UT-A Collected with 0 and 30 Degree Tilts

EMDB-16112: 
Human Urea Transporter UT-B/UT1 in Complex with Inhibitor UTBinh-14

PDB-8blo: 
Human Urea Transporter UT-A (N-Terminal Domain Model)

PDB-8blp: 
Human Urea Transporter UT-B/UT1 in Complex with Inhibitor UTBinh-14

EMDB-29433: 
Cryo-EM structure of the Cas13bt3-crRNA-target RNA ternary complex in activated state

PDB-8fti: 
Cryo-EM structure of the Cas13bt3-crRNA-target RNA ternary complex in activated state

EMDB-41153: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf

EMDB-41154: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf

PDB-8tcf: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
