-Search query
-Search result
Showing 1 - 50 of 74 items for (author: schmid & sl)

EMDB-43813: 
VIR-7229 Fab fragment bound the SARS-CoV-2 BA.2.86 spike trimer (local refinement of the BA 2.86 RBD/VIR-7229 VHVL)

EMDB-43842: 
VIR-7229 Fab fragment bound the BA.2.86 spike trimer (global refinement)

PDB-9asd: 
VIR-7229 Fab fragment bound the SARS-CoV-2 BA.2.86 spike trimer (local refinement of the BA 2.86 RBD/VIR-7229 VHVL)

PDB-9au2: 
VIR-7229 Fab fragment bound the BA.2.86 spike trimer (global refinement)

EMDB-18334: 
Cryo-EM structure of the inward-facing FLVCR1

EMDB-18335: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

EMDB-18336: 
Cryo-EM structure of the inward-facing FLVCR2

EMDB-18337: 
Cryo-EM structure of the outward-facing FLVCR2

EMDB-18339: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

EMDB-19009: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

EMDB-29530: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-29531: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-40240: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

PDB-8fxb: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

PDB-8fxc: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

PDB-8s9g: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-29207: 
CryoET tomogram of mitochondria in BACHD mouse model neuron

EMDB-29208: 
CryoET tomogram of BACHD mouse model neuron showing sheet aggregates

EMDB-29210: 
CryoET tomogram of purified mitochondria from HD patient iPSC-derived neuron (Q109)

EMDB-29211: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with PIAS1 hetKO treatment

EMDB-28668: 
CryoET tomogram of iPSC-derived control non-HD neuron (Q18)
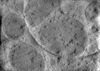
EMDB-28944: 
CryoET tomogram of iPSC-derived control non-HD neuron (Q20)

EMDB-28946: 
CryoET tomogram of HD patient iPSC-derived neuron (Q53)

EMDB-29074: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66)
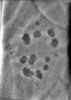
EMDB-29075: 
CryoET tomogram of HD patient iPSC-derived neuron (Q77)
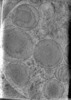
EMDB-29076: 
CryoET tomogram of HD patient iPSC-derived neuron (Q109)

EMDB-29079: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) showing sheet aggregate

EMDB-29080: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with PIAS1 hetKO treatment
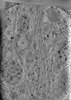
EMDB-29081: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with GRSF1 KD treatment

EMDB-29083: 
CryoET tomogram of mitochondria in WT mouse neuron

EMDB-29084: 
CryoET tomogram of mitochondria in BACHD-dN17 mouse model neuron

EMDB-28558: 
SARS-CoV-2 BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment (local refinement of the RBD and S2X324)

EMDB-28559: 
SARS-CoV-2 Omicron BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment

PDB-8erq: 
SARS-CoV-2 BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment (local refinement of the RBD and S2X324)

PDB-8err: 
SARS-CoV-2 Omicron BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment

EMDB-24894: 
Ab16 Fab in complex with SARS-CoV-2 Spike (6P)

EMDB-24895: 
Ab20 in complex with SARS-CoV-2 spike (6P)

EMDB-24347: 
SARS-CoV-2 S glycoprotein in complex with S2X259 Fab

EMDB-24365: 
SARS-CoV-2 S bound to S2X259 Fab (local refinement of the RBD/S2X259 variable domains)

PDB-7ra8: 
SARS-CoV-2 S glycoprotein in complex with S2X259 Fab

PDB-7ral: 
SARS-CoV-2 S bound to S2X259 Fab (local refinement of the RBD/S2X259 variable domains)

EMDB-23914: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain

EMDB-23915: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain

PDB-7mlz: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain

PDB-7mm0: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody B1-182.1 that targets the receptor-binding domain

EMDB-23498: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain

EMDB-23499: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain

PDB-7lrs: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain

PDB-7lrt: 
Cryo-EM structure of SARS-CoV-2 spike in complex with neutralizing antibody A23-58.1 that targets the receptor-binding domain

EMDB-24279: 
Cryo-EM map of SARS-CoV-2 HexaPro Spike in complex with Ab090 Fab
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
