-Search query
-Search result
Showing 1 - 50 of 260 items for (author: saunder & ko)
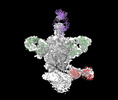
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.

EMDB-44740: 
HIV Envelope trimer CH505 SOSIP.664 in complex with three CH103 E75K/D76N mutant antibody Fabs

EMDB-44644: 
SARS CoV2 spike in complex with NTD-directed antibody Fab DH1052

EMDB-44736: 
HIV Envelope trimer BG505 SOSIP.664 in complex with wild type CH103 antibody

EMDB-44738: 
HIV Envelope BG505 SOSIP.664 in complex with one Fab of CH103 E75K/D76N mutant

EMDB-44739: 
HIV Envelope trimer CH505 SOSIP.664 in complex with wild type CH103 antibody

EMDB-40853: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab
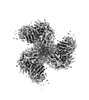
EMDB-40854: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab

PDB-8sxi: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab

PDB-8sxj: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab

EMDB-40856: 
Single particle reconstruction of the human LINE-1 ORF2p without substrate (apo)
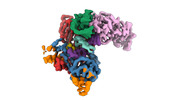
EMDB-40858: 
Structure of LINE-1 ORF2p with template:primer hybrid
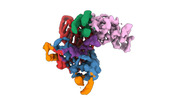
EMDB-40859: 
Structure of LINE-1 ORF2p with an oligo(A) template

PDB-8sxt: 
Structure of LINE-1 ORF2p with template:primer hybrid

PDB-8sxu: 
Structure of LINE-1 ORF2p with an oligo(A) template

EMDB-41810: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-41820: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41823: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41838: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to partially open HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

PDB-8u1d: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
EMDB-27706: 
Vaccine elicited Antibody MU89 bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
EMDB-27776: 
Vaccine elicited Antibody MU89+S27Y bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer

EMDB-40273: 
CryoEM structure of VRC01-CH848.0358.80

EMDB-40274: 
CryoEM structure of VRC01-CH848.0836.10

EMDB-40275: 
CryoEM structure of DH270.6-CH848.0526.25

EMDB-40277: 
CryoEM structure of DH270.6-CH848.10.17

EMDB-40278: 
CryoEM structure of DH270.5-CH848.10.17

EMDB-40279: 
CryoEM structure of VRC01-CH848.10.17

EMDB-40280: 
CryoEM structure of DH270.4-CH848.10.17

EMDB-40281: 
CryoEM structure of VRC01-CH848.0526.25

EMDB-40282: 
CryoEM structure of DH270.UCA.G57R-CH848.10.17DT

EMDB-40283: 
CryoEM structure of DH270.UCA-CH848.10.17DT

EMDB-40284: 
CryoEM structure of DH270.3-CH848.10.17

EMDB-40285: 
CryoEM structure of DH270.I5.6-CH848.10.17

EMDB-40286: 
CryoEM structure of DH270.I4.6-CH848.10.17

EMDB-40287: 
CryoEM structure of DH270.I3-CH848.10.17

EMDB-40288: 
CryoEM structure of DH270.I2-CH848.10.17

EMDB-40289: 
CryoEM structure of DH270.2-CH848.10.17

EMDB-40290: 
CryoEM structure of DH270.1-CH848.10.17

EMDB-40291: 
CryoEM structure of DH270.I1.6-CH848.10.17

PDB-8sal: 
CryoEM structure of VRC01-CH848.0358.80

PDB-8san: 
CryoEM structure of VRC01-CH848.0836.10

PDB-8saq: 
CryoEM structure of DH270.6-CH848.0526.25

PDB-8sar: 
CryoEM structure of DH270.6-CH848.10.17

PDB-8sas: 
CryoEM structure of DH270.5-CH848.10.17

PDB-8sat: 
CryoEM structure of VRC01-CH848.10.17

PDB-8sau: 
CryoEM structure of DH270.4-CH848.10.17

PDB-8sav: 
CryoEM structure of VRC01-CH848.0526.25

PDB-8saw: 
CryoEM structure of DH270.UCA.G57R-CH848.10.17DT
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
