-Search query
-Search result
Showing all 20 items for (author: rothman & j)

EMDB-71039: 
Subtomogram average of Munc13-1 C1-C2B-MUN-C2C F1176N/I1215N mutant 2D crystal between lipid bilayers
Method: subtomogram averaging / : Grushin K, Radhakrishnan A

EMDB-25737: 
Subtomogram average of the hexagonal assembly in Munc13-1 C1-C2B-MUN-C2C 2D crystal between lipid bilayers
Method: subtomogram averaging / : Grushin K, Sindelar CV

EMDB-25738: 
Subtomogram average of Munc13-1 C1-C2B-MUN-C2C trimer within 2D crystal between lipid bilayers.
Method: subtomogram averaging / : Grushin K, Sindelar CV

EMDB-25739: 
Composite map of Lateral Munc13-1 C1-C2B-MUN-C2C molecule
Method: subtomogram averaging / : Grushin K, Sindelar CV

EMDB-25740: 
Composite map of Upright Munc13-1 C1-C2B-MUN-C2C molecule spanning two lipid bilayers
Method: subtomogram averaging / : Grushin K, Sindelar CV

EMDB-25741: 
A composite 3D map of Munc13-1 C1-C2B-MUN-C2C 2D crystal between lipid bilayers.
Method: subtomogram averaging / : Grushin K, Sindelar CV

PDB-7t7c: 
The hexagonal organization of Munc13-1 C1-C2B-MUN-C2C domains between lipid bilayers
Method: subtomogram averaging / : Grushin K, Sindelar CV

PDB-7t7r: 
Structure of Munc13-1 C1-C2B-MUN-C2C trimer between lipid bilayers
Method: subtomogram averaging / : Grushin K, Sindelar CV

PDB-7t7v: 
Munc13-1 C1-C2B-MUN-C2C Lateral conformation on lipid bilayer surface
Method: subtomogram averaging / : Grushin K, Sindelar CV

PDB-7t7x: 
Munc13-1 C1-C2B-MUN-C2C Upright conformation spanning two lipid bilayers
Method: subtomogram averaging / : Grushin K, Sindelar CV

PDB-7t81: 
Model of Munc13-1 C1-C2B-MUN-C2C 2D crystal between lipid bilayers.
Method: subtomogram averaging / : Grushin K, Sindelar CV
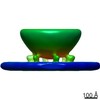
EMDB-23077: 
Hexameric protein arrangement under docked synaptic vesicles in primary hippocampal neurons
Method: subtomogram averaging / : Radhakrishnan A, Li X, Liu J

EMDB-9231: 
Cryo-EM structure of membrane-bound Synaptotagmin 1 C2AB domains in a complex with the SNAREpin assembly in presence of Mg2+
Method: helical / : Grushin K, Wang J, Coleman J, Rothman J, Sindelar C, Krishnakumar S

PDB-6mti: 
Synaptotagmin-1 C2A, C2B domains and SNARE-pin proteins (5CCI) individually docked into Cryo-EM map of C2AB-SNARE complexes helically organized on lipid nanotube surface in presence of Mg2+
Method: helical / : Grushin K, Wang J, Coleman J, Rothman J, Sindelar C, Krishnakumar S

EMDB-0413: 
Vesicle-plasma interface under the docked vesicles in NGF-differentiated PC12 cells without imposed symmetry
Method: subtomogram averaging / : Li X, Radhakrishnan A, Grushin K, Krishnakumar S, Liu J, Rothman J

EMDB-0414: 
Vesicle-plasma interface under the docked vesicles in NGF-differentiated PC12 cells with imposed 6-fold rotational symmetry
Method: subtomogram averaging / : Li X, Radhakrishnan A, Grushin K, Krishnakumar S, Liu J, Rothman J

EMDB-1472: 
Structure of bacteriophage N4 wild-type and mutants, determined by cryo-electron microscopy.
Method: single particle / : Choi KH, McPartland J, Kaganman I, Bowman VD, Rothman-Denes LB, Rossmann MG

EMDB-1475: 
Structure of bacteriophage N4
Method: single particle / : Choi KH, McPartland J, Kaganman I, Bowman VD, Rothman-Denes LB, Rossmann MG

EMDB-1476: 
Structure of a bacteriophage N4 mutant lacking gp65
Method: single particle / : Choi KH, McPartland J, Kaganman I, Bowman VD, Rothman-Denes LB, Rossmann MG

EMDB-1509: 
Structure of a bacteriophage N4 mutant lacking gp17
Method: single particle / : Choi KH, McPartland J, Kaganman I, Bowman VD, Rothman-Denes LB, Rossmann MG
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
