+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1475 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of bacteriophage N4 | |||||||||
 Map data Map data | This is an asymmetric reconstruction of bacteriophage N4 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Bacteriophage N4 / Podoviridae / non-contractile tail sheath / tail tube / appendages | |||||||||
| Biological species |  Enterobacteria phage N4 (virus) Enterobacteria phage N4 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 29.0 Å | |||||||||
 Authors Authors | Choi KH / McPartland J / Kaganman I / Bowman VD / Rothman-Denes LB / Rossmann MG | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2008 Journal: J Mol Biol / Year: 2008Title: Insight into DNA and protein transport in double-stranded DNA viruses: the structure of bacteriophage N4. Authors: Kyung H Choi / Jennifer McPartland / Irene Kaganman / Valorie D Bowman / Lucia B Rothman-Denes / Michael G Rossmann /  Abstract: Bacteriophage N4 encapsidates a 3500-aa-long DNA-dependent RNA polymerase (vRNAP), which is injected into the host along with the N4 genome upon infection. The three-dimensional structures of wild- ...Bacteriophage N4 encapsidates a 3500-aa-long DNA-dependent RNA polymerase (vRNAP), which is injected into the host along with the N4 genome upon infection. The three-dimensional structures of wild-type and mutant N4 viruses lacking gp17, gp50, or gp65 were determined by cryoelectron microscopy. The virion has an icosahedral capsid with T=9 quasi-symmetry that encapsidates well-organized double-stranded DNA and vRNAP. The tail, attached at a unique pentameric vertex of the head, consists of a neck, 12 appendages, and six ribbons that constitute a non-contractile sheath around a central tail tube. Comparison of wild-type and mutant virus structures in conjunction with bioinformatics established the identity and virion locations of the major capsid protein (gp56), a decorating protein (gp17), the vRNAP (gp50), the tail sheath (gp65), the appendages (gp66), and the portal protein (gp59). The N4 virion organization provides insight into its assembly and suggests a mechanism for genome and vRNAP transport strategies utilized by this unique system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1475.map.gz emd_1475.map.gz | 205 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1475-v30.xml emd-1475-v30.xml emd-1475.xml emd-1475.xml | 7.2 KB 7.2 KB | Display Display |  EMDB header EMDB header |
| Images |  1475.gif 1475.gif | 92.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1475 http://ftp.pdbj.org/pub/emdb/structures/EMD-1475 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1475 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1475 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1475.map.gz / Format: CCP4 / Size: 276 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1475.map.gz / Format: CCP4 / Size: 276 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is an asymmetric reconstruction of bacteriophage N4 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.54 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
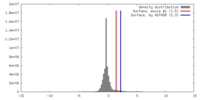
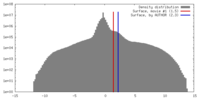
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bacteriophage N4
| Entire | Name:  Bacteriophage N4 (virus) Bacteriophage N4 (virus) |
|---|---|
| Components |
|
-Supramolecule #1000: Bacteriophage N4
| Supramolecule | Name: Bacteriophage N4 / type: sample / ID: 1000 / Number unique components: 10 |
|---|
-Supramolecule #1: Enterobacteria phage N4
| Supramolecule | Name: Enterobacteria phage N4 / type: virus / ID: 1 / Name.synonym: bacteriophage N4 / NCBI-ID: 10752 / Sci species name: Enterobacteria phage N4 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: bacteriophage N4 |
|---|---|
| Host (natural) | synonym: BACTERIA(EUBACTERIA) |
| Virus shell | Shell ID: 1 / Diameter: 740 Å / T number (triangulation number): 9 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG |
|---|---|
| Image recording | Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 14 µm |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.75 µm / Nominal defocus min: 1.13 µm / Nominal magnification: 39500 |
| Sample stage | Specimen holder: eucentric / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 29.0 Å / Resolution method: FSC 0.5 CUT-OFF / Number images used: 2489 |
|---|
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)