-Search query
-Search result
Showing 1 - 50 of 1,004 items for (author: phan & n)

EMDB-45001: 
Structure of Mnx H340A complex from Bacillus sp. PL-12

PDB-9bxa: 
Structure of Mnx H340A complex from Bacillus sp. PL-12

EMDB-43551: 
CCHFV GP38 bound with ADI-46143 and ADI-46158 Fabs

EMDB-43552: 
CCHFV GP38 bound with ADI-58062 and ADI-63530 Fabs

EMDB-43553: 
CCHFV GP38 bound with ADI-58026 and ADI-63547 Fabs

EMDB-43604: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs

PDB-8vww: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs

EMDB-43296: 
Constituent EM map: Focused refinement S100A1 of mouse RyR1 in complex with S100A1 (EGTA-only dataset)

EMDB-44351: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, V1

PDB-9b8p: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, V1

EMDB-44350: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, Vo

EMDB-44352: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, peripheral stalks

EMDB-44353: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3

EMDB-44354: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 2

EMDB-44355: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 1

PDB-9b8o: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, Vo

PDB-9b8q: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, peripheral stalks

PDB-9brb: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 1

PDB-9brc: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 2

PDB-9brd: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3

EMDB-18878: 
Cryo-EM structure of the Asgard archaeal Argonaute HrAgo1 bound to a guide RNA

PDB-8r3z: 
Cryo-EM structure of the Asgard archaeal Argonaute HrAgo1 bound to a guide RNA

EMDB-43592: 
PDI-containing spoke of a hexagonal wireframe DNA origami

EMDB-16543: 
Structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-His(5,Ser)

EMDB-16544: 
Structure of human mitochondrial RNase Z in complex with mitochondrial pre-tRNA-His(0,Ser)

EMDB-16545: 
Structure of human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNA-Ile
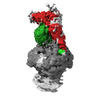
EMDB-16547: 
Structure of human mitochondrial MRPP1-MRPP2 in complex with mitochondrial pre-tRNA-Ile

EMDB-16667: 
Consensus Refinement - Map 1

EMDB-16668: 
Local refinement - Map 2

EMDB-16669: 
Cryo-EM structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-His(5,Ser)

EMDB-16673: 
Consensus Refinement - Map 1

EMDB-16674: 
Local refinement - Map 2 (mask 1)

EMDB-16675: 
Local refinement - map 3 (Mask 2)

EMDB-16690: 
Consensus Refinement - Map 1

EMDB-16691: 
Local refinement - Map 2 (Mask 1)

EMDB-16692: 
Global Refinement - Map 1

PDB-8cbk: 
Structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-His(5,Ser)

PDB-8cbl: 
Structure of human mitochondrial RNase Z in complex with mitochondrial pre-tRNA-His(0,Ser)

PDB-8cbm: 
Structure of human mitochondrial CCA-adding enzyme in complex with mitochondrial pre-tRNA-Ile

PDB-8cbo: 
Structure of human mitochondrial MRPP1-MRPP2 in complex with mitochondrial pre-tRNA-Ile

EMDB-29330: 
N332-GT5 SOSIP in complex with base polyclonal Fabs isolated at day 42 from protein immunized wild type mice

EMDB-29333: 
N332-GT5 SOSIP in complex with base polyclonal Fabs isolated at day 42 from protein immunized BG18HCgl knock-in mice

EMDB-29334: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 16 from mRNA immunized wild type mice

EMDB-29335: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 42 from mRNA immunized BG18HCgl knock-in mice

EMDB-43191: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 14 from N332-GT2 nanoparticle-immunized BG18HCgl knock-in mice

EMDB-43192: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 15 from N332-GT5 nanoparticle-immunized BG18HCgl knock-in mice
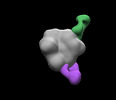
EMDB-28937: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 42 from protein immunized wild type mice
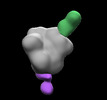
EMDB-28938: 
N332-GT5 SOSIP in complex with V1V3 and base polyclonal Fabs isolated at day 16 from protein immunized BG18HCgl knock-in mice
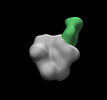
EMDB-28939: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 42 from mRNA immunized wild type mice
EMDB-28940: 
N332-GT5 SOSIP in complex with V1V3 polyclonal Fabs isolated at day 16 from mRNA immunized BG18HCgl knock-in mice
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
