-Search query
-Search result
Showing 1 - 50 of 120 items for (author: herrmann & e)

EMDB-42150: 
Human Mitochondrial DNA Polymerase Gamma Binary Complex

PDB-8udl: 
Human Mitochondrial DNA Polymerase Gamma Binary Complex

EMDB-42343: 
The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2+ (class 5 reconstruction)

EMDB-42344: 
The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2+ (class 6 reconstruction)

PDB-8uk2: 
The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2+ (class 5 reconstruction)

PDB-8uk3: 
The rotavirus VP5*/VP8* conformational transition permeabilizes membranes to Ca2+ (class 6 reconstruction)

EMDB-18698: 
Structural insights into the activation mechanism of antimicrobial GBP1: Polymeric assembly of GBP1

EMDB-18806: 
Structural insights into the activation mechanism of antimicrobial GBP1: Membrane-bound GBP1 oligomer

PDB-8r1a: 
Model of the membrane-bound GBP1 oligomer

EMDB-27171: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer (local refinement of subunit B)

EMDB-27154: 
Human mitochondrial DNA polymerase gamma ternary complex with GC basepair

EMDB-27155: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in replication conformer

EMDB-27163: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in intermediate conformer

EMDB-27169: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer (consensus)

EMDB-27170: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer (local refinement of subunit A and primer/template DNA)

EMDB-27172: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer (composite)

PDB-8d33: 
Human mitochondrial DNA polymerase gamma ternary complex with GC basepair

PDB-8d37: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in replication conformer

PDB-8d3r: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in intermediate conformer

PDB-8d42: 
Human mitochondrial DNA polymerase gamma ternary complex with GT basepair in editing conformer (composite)

EMDB-16457: 
Drosophila melanogaster Rab7 GEF complex Mon1-Ccz1-Bulli

EMDB-16462: 
MCB complex from drosophila melanogaster, local refinement map of the bottom part of the complex

EMDB-16463: 
MCB complex from drosophila melanogaster, local refinement map of the MCB core

EMDB-16464: 
MCB complex from drosophila melanogaster, consensus refinement

PDB-8c7g: 
Drosophila melanogaster Rab7 GEF complex Mon1-Ccz1-Bulli

EMDB-16128: 
Tomogram of a late endosome of A549 cell infected with influenza A virus.

EMDB-16129: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6C).

EMDB-16130: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6E)

EMDB-16131: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6G)

EMDB-16132: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6I,K,M)

EMDB-16133: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6O)

EMDB-15905: 
Cryo-EM structure of the E.coli 70S ribosome in complex with the antibiotic Myxovalargin B.

PDB-8b7y: 
Cryo-EM structure of the E.coli 70S ribosome in complex with the antibiotic Myxovalargin B.

EMDB-14121: 
Cryo-EM structure of the E.coli 50S ribosomal subunit in complex with the antibiotic Myxovalargin A.

PDB-7qq3: 
Cryo-EM structure of the E.coli 50S ribosomal subunit in complex with the antibiotic Myxovalargin A.

EMDB-15362: 
Cryo-EM structure of Darobactin 22 bound BAM complex

EMDB-15363: 
Cryo-EM structure of Darobactin 9 bound BAM complex

PDB-8adg: 
Cryo-EM structure of Darobactin 22 bound BAM complex

PDB-8adi: 
Cryo-EM structure of Darobactin 9 bound BAM complex

EMDB-15705: 
Tomogram of a late endosome of A549 cell (Figure 1R)

EMDB-15707: 
Tomogram of a late endosome of A549 cell treated with IFN-beta (Figure 1T)
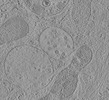
EMDB-15708: 
Tomogram of a late endosome of A549-IFITM3 cell (Figure 1V)

EMDB-15131: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S7)

EMDB-15130: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure 4E)
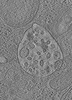
EMDB-15132: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S8)

EMDB-15133: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S9)

EMDB-14066: 
Structure of the Rab GEF complex Mon1-Ccz1

PDB-7qla: 
Structure of the Rab GEF complex Mon1-Ccz1

EMDB-21955: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Upright conformation

EMDB-21956: 
Cryo-EM reconstruction of VP5*/VP8* assembly from rhesus rotavirus particles - Intermediate conformation
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
