+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Mitochondrial DNA Polymerase Gamma Binary Complex | |||||||||
 Map data Map data | LocSpiral Processed Map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mitochondrial / DNA Polymerase / TRANSFERASE / TRANSFERASE-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationgamma DNA polymerase complex / positive regulation of DNA-directed DNA polymerase activity / mitochondrial chromosome / Strand-asynchronous mitochondrial DNA replication / mitochondrial DNA replication / DNA replication proofreading / single-stranded DNA 3'-5' DNA exonuclease activity / Hydrolases; Acting on ester bonds; Exodeoxyribonucleases producing 5'-phosphomonoesters / DNA metabolic process / DNA polymerase processivity factor activity ...gamma DNA polymerase complex / positive regulation of DNA-directed DNA polymerase activity / mitochondrial chromosome / Strand-asynchronous mitochondrial DNA replication / mitochondrial DNA replication / DNA replication proofreading / single-stranded DNA 3'-5' DNA exonuclease activity / Hydrolases; Acting on ester bonds; Exodeoxyribonucleases producing 5'-phosphomonoesters / DNA metabolic process / DNA polymerase processivity factor activity / Lyases; Carbon-oxygen lyases; Other carbon-oxygen lyases / mitochondrial nucleoid / 5'-deoxyribose-5-phosphate lyase activity / base-excision repair, gap-filling / DNA polymerase binding / 3'-5' exonuclease activity / mitochondrion organization / Transcriptional activation of mitochondrial biogenesis / base-excision repair / DNA-templated DNA replication / protease binding / double-stranded DNA binding / DNA-directed DNA polymerase / in utero embryonic development / DNA-directed DNA polymerase activity / mitochondrial matrix / chromatin binding / protein-containing complex / mitochondrion / DNA binding / identical protein binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / synthetic construct (others) Homo sapiens (human) / synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.37 Å | |||||||||
 Authors Authors | Park J / Yin YW | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2024 Journal: Sci Adv / Year: 2024Title: An interaction network in the polymerase active site is a prerequisite for Watson-Crick base pairing in Pol γ. Authors: Joon Park / Geoffrey K Herrmann / Arkanil Roy / Christie K Shumate / G Andrés Cisneros / Y Whitney Yin /  Abstract: The replication accuracy of DNA polymerase gamma (Pol γ) is essential for mitochondrial genome integrity. Mutation of human Pol γ arginine-853 has been linked to neurological diseases. Although not ...The replication accuracy of DNA polymerase gamma (Pol γ) is essential for mitochondrial genome integrity. Mutation of human Pol γ arginine-853 has been linked to neurological diseases. Although not a catalytic residue, Pol γ arginine-853 mutants are void of polymerase activity. To identify the structural basis for the disease, we determined a crystal structure of the Pol γ mutant ternary complex with correct incoming nucleotide 2'-deoxycytidine 5'-triphosphate (dCTP). Opposite to the wild type that undergoes open-to-closed conformational changes when bound to a correct nucleotide that is essential for forming a catalytically competent active site, the mutant complex failed to undergo the conformational change, and the dCTP did not base pair with its Watson-Crick complementary templating residue. Our studies revealed that arginine-853 coordinates an interaction network that aligns the 3'-end of primer and dCTP with the catalytic residues. Disruption of the network precludes the formation of Watson-Crick base pairing and closing of the active site, resulting in an inactive polymerase. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42150.map.gz emd_42150.map.gz | 64.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42150-v30.xml emd-42150-v30.xml emd-42150.xml emd-42150.xml | 23.7 KB 23.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_42150.png emd_42150.png | 128.2 KB | ||
| Filedesc metadata |  emd-42150.cif.gz emd-42150.cif.gz | 7.5 KB | ||
| Others |  emd_42150_additional_1.map.gz emd_42150_additional_1.map.gz emd_42150_half_map_1.map.gz emd_42150_half_map_1.map.gz emd_42150_half_map_2.map.gz emd_42150_half_map_2.map.gz | 63 MB 115.9 MB 115.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42150 http://ftp.pdbj.org/pub/emdb/structures/EMD-42150 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42150 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42150 | HTTPS FTP |
-Related structure data
| Related structure data |  8udlMC  8udkC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42150.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42150.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | LocSpiral Processed Map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.86 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: Unprocessed Map
| File | emd_42150_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unprocessed Map | ||||||||||||
| Projections & Slices |
| ||||||||||||
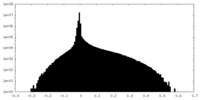
| Density Histograms |
-Half map: Half Map A
| File | emd_42150_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
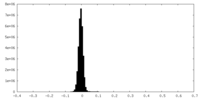
| Density Histograms |
-Half map: Half Map B
| File | emd_42150_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half Map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human Mitochondrial DNA Polymerase Gamma Binary Complex
| Entire | Name: Human Mitochondrial DNA Polymerase Gamma Binary Complex |
|---|---|
| Components |
|
-Supramolecule #1: Human Mitochondrial DNA Polymerase Gamma Binary Complex
| Supramolecule | Name: Human Mitochondrial DNA Polymerase Gamma Binary Complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: DNA polymerase subunit gamma-1,DNA polymerase subunit gamma-2
| Supramolecule | Name: DNA polymerase subunit gamma-1,DNA polymerase subunit gamma-2 type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: DNA
| Supramolecule | Name: DNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #3-#4 |
|---|
-Macromolecule #1: DNA polymerase subunit gamma-1
| Macromolecule | Name: DNA polymerase subunit gamma-1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 139.730703 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: MSRLLWRKVA GATVGPGPVP APGRWVSSSV PASDPSDGQR RRQQQQQQQQ QQQQQPQQPQ VLSSEGGQLR HNPLDIQMLS RGLHEQIFG QGGEMPGEAA VRRSVEHLQK HGLWGQPAVP LPDVELRLPP LYGDNLDQHF RLLAQKQSLP YLEAANLLLQ A QLPPKPPA ...String: MSRLLWRKVA GATVGPGPVP APGRWVSSSV PASDPSDGQR RRQQQQQQQQ QQQQQPQQPQ VLSSEGGQLR HNPLDIQMLS RGLHEQIFG QGGEMPGEAA VRRSVEHLQK HGLWGQPAVP LPDVELRLPP LYGDNLDQHF RLLAQKQSLP YLEAANLLLQ A QLPPKPPA WAWAEGWTRY GPEGEAVPVA IPEERALVFD VEVCLAEGTC PTLAVAISPS AWYSWCSQRL VEERYSWTSQ LS PADLIPL EVPTGASSPT QRDWQEQLVV GHNVSFDRAH IREQYLIQGS RMRFLDTMSM HMAISGLSSF QRSLWIAAKQ GKH KVQPPT KQGQKSQRKA RRGPAISSWD WLDISSVNSL AEVHRLYVGG PPLEKEPREL FVKGTMKDIR ENFQDLMQYC AQDV WATHE VFQQQLPLFL ERCPHPVTLA GMLEMGVSYL PVNQNWERYL AEAQGTYEEL QREMKKSLMD LANDACQLLS GERYK EDPW LWDLEWDLQE FKQKKAKKVK KEPATASKLP IEGAGAPGDP MDQEDLGPCS EEEEFQQDVM ARACLQKLKG TTELLP KRP QHLPGHPGWY RKLCPRLDDP AWTPGPSLLS LQMRVTPKLM ALTWDGFPLH YSERHGWGYL VPGRRDNLAK LPTGTTL ES AGVVCPYRAI ESLYRKHCLE QGKQQLMPQE AGLAEEFLLT DNSAIWQTVE ELDYLEVEAE AKMENLRAAV PGQPLALT A RGGPKDTQPS YHHGNGPYND VDIPGCWFFK LPHKDGNSCN VGSPFAKDFL PKMEDGTLQA GPGGASGPRA LEINKMISF WRNAHKRISS QMVVWLPRSA LPRAVIRHPD YDEEGLYGAI LPQVVTAGTI TRRAVEPTWL TASNARPDRV GSELKAMVQA PPGYTLVGA DVDSQELWIA AVLGDAHFAG MHGCTAFGWM TLQGRKSRGT DLHSKTATTV GISREHAKIF NYGRIYGAGQ P FAERLLMQ FNHRLTQQEA AEKAQQMYAA TKGLRWYRLS DEGEWLVREL NLPVDRTEGG WISLQDLRKV QRETARKSQW KK WEVVAER AWKGGTESEM FNKLESIATS DIPRTPVLGC CISRALEPSA VQEEFMTSRV NWVVQSSAVD YLHLMLVAMK WLF EEFAID GRFCISIHDE VRYLVREEDR YRAALALQIT NLLTRCMFAY KLGLNDLPQS VAFFSAVDID RCLRKEVTMD CKTP SNPTG MERRYGIPQG EALDIYQIIE LTKGSLEKRS QPGP UniProtKB: DNA polymerase subunit gamma-1 |
-Macromolecule #2: DNA polymerase subunit gamma-2, mitochondrial
| Macromolecule | Name: DNA polymerase subunit gamma-2, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 54.991 KDa |
| Recombinant expression | Organism: Insect cell expression vector pTIE1 (others) |
| Sequence | String: MRSRVAVRAC HKVCRCLLSG FGGRVDAGQP ELLTERSSPK GGHVKSHAEL EGNGEHPEAP GSGEGSEALL EICQRRHFLS GSKQQLSRD SLLSGCHPGF GPLGVELRKN LAAEWWTSVV VFREQVFPVD ALHHKPGPLL PGDSAFRLVS AETLREILQD K ELSKEQLV ...String: MRSRVAVRAC HKVCRCLLSG FGGRVDAGQP ELLTERSSPK GGHVKSHAEL EGNGEHPEAP GSGEGSEALL EICQRRHFLS GSKQQLSRD SLLSGCHPGF GPLGVELRKN LAAEWWTSVV VFREQVFPVD ALHHKPGPLL PGDSAFRLVS AETLREILQD K ELSKEQLV AFLENVLKTS GKLRENLLHG ALEHYVNCLD LVNKRLPYGL AQIGVCFHPV FDTKQIRNGV KSIGEKTEAS LV WFTPPRT SNQWLDFWLR HRLQWWRKFA MSPSNFSSSD CQDEEGRKGN KLYYNFPWGK ELIETLWNLG DHELLHMYPG NVS KLHGRD GRKNVVPCVL SVNGDLDRGM LAYLYDSFQL TENSFTRKKN LHRKVLKLHP CLAPIKVALD VGRGPTLELR QVCQ GLFNE LLENGISVWP GYLETMQSSL EQLYSKYDEM SILFTVLVTE TTLENGLIHL RSRDTTMKEM MHISKLKDFL IKYIS SAKN V UniProtKB: DNA polymerase subunit gamma-2 |
-Macromolecule #3: DNA (5'-D(P*AP*AP*AP*AP*CP*GP*AP*CP*GP*GP*CP*CP*AP*GP*TP*GP*CP*CP...
| Macromolecule | Name: DNA (5'-D(P*AP*AP*AP*AP*CP*GP*AP*CP*GP*GP*CP*CP*AP*GP*TP*GP*CP*CP*AP*TP*AP*C)-3') type: dna / ID: 3 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 6.739384 KDa |
| Sequence | String: (DA)(DA)(DA)(DA)(DC)(DG)(DA)(DC)(DG)(DG) (DC)(DC)(DA)(DG)(DT)(DG)(DC)(DC)(DA)(DT) (DA)(DC) |
-Macromolecule #4: DNA (5'-D(P*AP*GP*GP*TP*AP*TP*GP*GP*CP*AP*CP*TP*GP*GP*CP*CP*GP*TP...
| Macromolecule | Name: DNA (5'-D(P*AP*GP*GP*TP*AP*TP*GP*GP*CP*AP*CP*TP*GP*GP*CP*CP*GP*TP*CP*GP*TP*TP*TP*T)-3') type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 7.407761 KDa |
| Sequence | String: (DA)(DG)(DG)(DT)(DA)(DT)(DG)(DG)(DC)(DA) (DC)(DT)(DG)(DG)(DC)(DC)(DG)(DT)(DC)(DG) (DT)(DT)(DT)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)