-Search query
-Search result
Showing all 48 items for (author: garaeva & n)
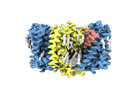
EMDB-18621: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)
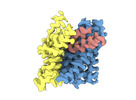
EMDB-18622: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)
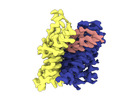
EMDB-18623: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)
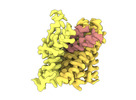
EMDB-18624: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)
EMDB-18625: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)
EMDB-18626: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)
EMDB-18627: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)
EMDB-18628: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)

PDB-8qro: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)

PDB-8qrp: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)

PDB-8qrq: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)

PDB-8qrr: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)

PDB-8qrs: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)

PDB-8qru: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)

PDB-8qrv: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)

PDB-8qrw: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)

EMDB-16334: 
Cryo-EM structure of a Staphylococus aureus 30S-RbfA complex

PDB-8byv: 
Cryo-EM structure of a Staphylococus aureus 30S-RbfA complex

EMDB-16045: 
The complex of the immature 30S ribosomal subunit with RimP from S. aureus

EMDB-16046: 
The complex of the immature 30S ribosomal subunit with RimP from S. aureus (body)

EMDB-16048: 
Mature 30S ribosomal subunit from Staphylococcus aureus

EMDB-16049: 
The complex of the immature 30S ribosomal subunit with RimP from S. aureus (composite map)

PDB-8bh6: 
Mature 30S ribosomal subunit from Staphylococcus aureus

PDB-8bh7: 
The complex of immature 30S ribosomal subunit with Ribosome maturation factor P (RimP) from Staphylococcus aureus

EMDB-12142: 
ASCT2 in the presence of the inhibitor Lc-BPE in the outward-open conformation.

EMDB-12143: 
ASCT2 in the presence of the inhibitor ERA-21 in the outward-open conformation.

PDB-7bcq: 
ASCT2 in the presence of the inhibitor Lc-BPE (position "up") in the outward-open conformation.

PDB-7bcs: 
ASCT2 in the presence of the inhibitor Lc-BPE (position "down") in the outward-open conformation.

PDB-7bct: 
ASCT2 in the presence of the inhibitor ERA-21 in the outward-open conformation.

EMDB-12314: 
Structure of glutamate transporter homologue in complex with Sybody

PDB-7ngh: 
Structure of glutamate transporter homologue in complex with Sybody

PDB-7p77: 
SARS-CoV-2 spike protein in complex with sybody#15 and sybody#68 in a 3up conformation

PDB-7p78: 
SARS-CoV-2 spike protein in complex with sybody#15 and sybody#68 in a 1up/1up-out/1down conformation

PDB-7p79: 
SARS-CoV-2 spike protein in complex with sybodyb#15 in a 1up/1up-out/1down conformation.

PDB-7p7a: 
SARS-CoV-2 spike protein in complex with sybody#68 in a 2up/1flexible conformation

PDB-7p7b: 
SARS-CoV-2 spike protein in complex with sybody no68 in a 1up/2down conformation

EMDB-12082: 
Cryo-EM map of the SARS-CoV-2 spike protein in complex with sybody#15 and sybody#68 in a 3up conformation.

EMDB-12083: 
Cryo-EM map of the SARS-CoV-2 spike protein in complex with sybody#15 and sybody#68 in a 1up/1up-out/1down conformation.

EMDB-12084: 
Cryo-EM map of the SARS-CoV-2 spike protein in complex with sybody#15 in a 1up/1up-out/1down conformation.

EMDB-12085: 
Cryo-EM map of the SARS-CoV-2 spike protein in complex with sybody#68 in a 2up/1flexible conformation.

EMDB-12086: 
Cryo-EM map of the SARS-CoV-2 spike protein in complex with sybody#68 in a 1up/2down conformation.

EMDB-10016: 
Inward-open structure of the ASCT2 (SLC1A5) mutant C467R in presence of TBOA

EMDB-10017: 
Inward-open structure of the ASCT2 (SLC1A5) in absence of substrate

EMDB-10018: 
Inward-open structure of the ASCT2 (SLC1A5) mutant C467R in absence of substrate

PDB-6rvx: 
Inward-open structure of the ASCT2 (SLC1A5) mutant C467R in presence of TBOA

PDB-6rvy: 
Inward-open structure of the ASCT2 (SLC1A5) mutant C467R in absence of substrate

EMDB-4386: 
cryo-EM structure of the human neutral amino acid transporter ASCT2

PDB-6gct: 
cryo-EM structure of the human neutral amino acid transporter ASCT2
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
