-Search query
-Search result
Showing 1 - 50 of 74 items for (author: elias & p)
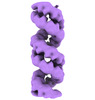
EMDB-19599: 
Structural characterization of Thogoto Virus nucleoprotein provides insights into RNA encapsidation and assembly

PDB-8ryt: 
Structural characterization of Thogoto Virus nucleoprotein provides insights into RNA encapsidation and assembly

EMDB-19837: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch

EMDB-19838: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site with 1-bp DNA mismatch

EMDB-19839: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch consensus map
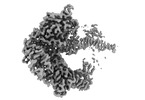
EMDB-19840: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch catalytic core focused map

EMDB-19841: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch processivity factor focused refinement

PDB-9enp: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state with 1-bp DNA mismatch

PDB-9enq: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site with 1-bp DNA mismatch

EMDB-17013: 
HSV-1 DNA polymerase-processivity factor complex in halted elongation state consensus map
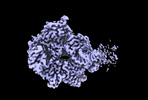
EMDB-17014: 
Consensus map of HSV-1 DNA polymerase-processivity factor complex in pre-translocation state

EMDB-17018: 
Consensus map of HSV-1 DNA polymerase-processivity factor complex in exonuclease state

EMDB-18214: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex - hexameric assembly
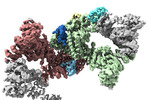
EMDB-18216: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused cullin dimer
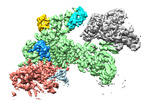
EMDB-18217: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused on E2-like density

EMDB-18218: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused dimeric core
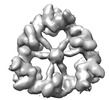
EMDB-18220: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 CPH domain
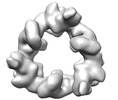
EMDB-18221: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 DOC domain
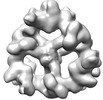
EMDB-18222: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 ARM9 domain
EMDB-18223: 
Structure of the hexameric CUL9-RBX1 complex with deletion of CUL9 ARIH-RBR element

EMDB-19179: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated conformation - symmetry expanded unneddylated dimer

PDB-8q7e: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex - hexameric assembly

PDB-8q7h: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated and neddylated conformation - focused cullin dimer

PDB-8rhz: 
Structure of CUL9-RBX1 ubiquitin E3 ligase complex in unneddylated conformation - symmetry expanded unneddylated dimer

EMDB-16918: 
Focused refinement map of HSV-1 DNA polymerase in pre-translocation state

EMDB-16919: 
Focused refinement map of HSV-1 DNA polymerase processivity factor in pre-translocation state
EMDB-16924: 
Focused refinement of HSV-1 DNA polymerase in halted elongation state

EMDB-16925: 
Focused refinement of HSV-1 DNA polymerase processivity factor in halted elongation state

EMDB-16927: 
Focused refinement map of HSV-1 DNA polymerase in exonuclease state

EMDB-16928: 
Focused refinement map of HSV-1 DNA polymerase processivity factor in exonuclease state

EMDB-16906: 
HSV-1 DNA polymerase-processivity factor complex in pre-translocation state

EMDB-16907: 
HSV-1 DNA polymerase-processivity factor complex in halted elongation state

EMDB-16909: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state

EMDB-16910: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site
EMDB-16911: 
HSV-1 DNA polymerase active site in alternative exonuclease state
EMDB-16912: 
HSV-1 DNA polymerase beta-hairpin loop

PDB-8oj6: 
HSV-1 DNA polymerase-processivity factor complex in pre-translocation state

PDB-8oj7: 
HSV-1 DNA polymerase-processivity factor complex in halted elongation state

PDB-8oja: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state

PDB-8ojb: 
HSV-1 DNA polymerase-processivity factor complex in exonuclease state active site

PDB-8ojc: 
HSV-1 DNA polymerase active site in alternative exonuclease state

PDB-8ojd: 
HSV-1 DNA polymerase beta-hairpin loop

EMDB-27382: 
Helical rods of far-red light-absorbing allophycocyanin in Synechococcus sp.

PDB-8ddy: 
Helical rods of far-red light-absorbing allophycocyanin in Synechococcus sp.

EMDB-35060: 
Clr4-H3K9 Nucleosome complex

EMDB-12142: 
ASCT2 in the presence of the inhibitor Lc-BPE in the outward-open conformation.

EMDB-12143: 
ASCT2 in the presence of the inhibitor ERA-21 in the outward-open conformation.

PDB-7bcq: 
ASCT2 in the presence of the inhibitor Lc-BPE (position "up") in the outward-open conformation.

PDB-7bcs: 
ASCT2 in the presence of the inhibitor Lc-BPE (position "down") in the outward-open conformation.

PDB-7bct: 
ASCT2 in the presence of the inhibitor ERA-21 in the outward-open conformation.
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
