-Search query
-Search result
Showing 1 - 50 of 759 items for (author: cui & t)

EMDB-41766: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant, scFv16, and dopamine

EMDB-41776: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant and dopamine
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
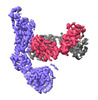
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
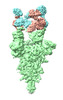
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
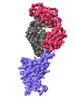
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
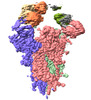
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535

EMDB-45364: 
HIV-1 intasome core bound with DTG

PDB-9c9m: 
HIV-1 intasome core bound with DTG

EMDB-37441: 
FCP tetramer in Chaetoceros gracilis

EMDB-37442: 
FCP pentamer in Chaetoceros gracilis

PDB-8wck: 
FCP tetramer in Chaetoceros gracilis

PDB-8wcl: 
FCP pentamer in Chaetoceros gracilis

EMDB-39645: 
The structure of HKU1-B S protein with bsAb1

EMDB-39646: 
the complex structure of the H4B6 Fab with the RBD of Omicron BA.5 S protein

PDB-8yww: 
The structure of HKU1-B S protein with bsAb1

PDB-8ywx: 
the complex structure of the H4B6 Fab with the RBD of Omicron BA.5 S protein

EMDB-38617: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab

EMDB-38618: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2

EMDB-38619: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)

EMDB-38620: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2

EMDB-38621: 
SARS-CoV-2 spike + IMCAS-123

EMDB-38823: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex

PDB-8xse: 
SARS-CoV-2 RBD + IMCAS-123 + IMCAS-72 Fab

PDB-8xsf: 
SARS-CoV-2 RBD + IMCAS-364 + hACE2

PDB-8xsi: 
SARS-CoV-2 RBD + IMCAS-364 (Local Refinement)

PDB-8xsj: 
SARS-CoV-2 Omicron BA.4 RBD + IMCAS-316 + ACE2

PDB-8xsl: 
SARS-CoV-2 spike + IMCAS-123

PDB-8y0y: 
Cryo-EM structure of the 123-316 scDb/PT-RBD complex
EMDB-16384: 
Tetrameric 5-HT3A receptor in Salipro (apo, asymmetric)
EMDB-16385: 
Tetrameric 5-HT3aR in Salipro (apo state, symmetric)
EMDB-16386: 
Tetrameric 5-HT3aR in Salipro (holo state, symmetric)
EMDB-16387: 
Tetrameric 5-HT3A receptor in Salipro (holo, asymmetric)

EMDB-36466: 
FCP trimer in diatom Thalassiosira pseudonana

PDB-8jp3: 
FCP trimer in diatom Thalassiosira pseudonana
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
