-Search query
-Search result
Showing 1 - 50 of 335 items for (author: chen & sj)

EMDB-37249: 
Cryo-EM structure of EBV gH/gL-gp42 in complex with fab 2C1

EMDB-41409: 
Cryo-EM structure of PCSK9 mimic HIT01-K21Q-R218E with AMG145 Fab
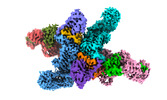
EMDB-37736: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 1)
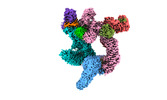
EMDB-37737: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 2)

EMDB-37739: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CDK5R1
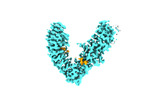
EMDB-37740: 
Local refinement of FEM1B bound with the C-degron of CCC89
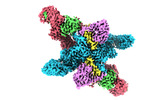
EMDB-37742: 
Cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CUX1 (conformation 1)
EMDB-37743: 
cryo-EM structure of CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CUX1 (conformation 2)
EMDB-37744: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 1)
EMDB-37745: 
cryo-EM structure of neddylated CUL2-RBX1-ELOB-ELOC-FEM1B bound with the C-degron of CCDC89 (conformation 2)

EMDB-37746: 
Local refinement of FEM1B bound with the C-degron of CUX1

EMDB-40650: 
Phosphoinositide phosphate 3 kinase gamma

EMDB-40651: 
Phosphoinositide phosphate 3 kinase gamma bound with ATP

EMDB-40652: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP

EMDB-40653: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and Gbetagamma

EMDB-40654: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 1

EMDB-40655: 
Phosphoinositide phosphate 3 kinase gamma bound with ADP and two Gbetagamma subunits in State 2

EMDB-41423: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40

EMDB-41424: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1

EMDB-41425: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2

EMDB-41777: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40

EMDB-41778: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1

EMDB-41779: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2
EMDB-28198: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with LLNL-199

EMDB-28199: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112
EMDB-41048: 
Lassa GPC Trimer in complex with Fab 8.11G and nanobody D5

EMDB-29696: 
In situ structure of GspD-beta-GspS complex on the bacterial outer membrane

EMDB-29697: 
In situ structure of GspD-beta-GspS complex on the bacterial outer membrane when GspD-beta is overexpressed solely

EMDB-29698: 
In situ structure of GspD-beta on the bacterial inner membrane

EMDB-29702: 
In situ structure of GspD-alpha on the bacterial inner membrane

EMDB-29703: 
In situ structure of GspD-alpha on the bacterial outer membrane

EMDB-36020: 
Cryo-EM structure of the Arabidopsis thaliana photosystem I(PSI-LHCII-ST2)

EMDB-36021: 
Cryo-EM structure of the Arabidopsis thaliana photosystem I(PSI-LHCII-ST2)

EMDB-36022: 
LHCI of PSI-LHCII from Arabidopsis thaliana(state2)

EMDB-36023: 
LHCII of PSI-LHCII from Arabidopsis thaliana(state2)

EMDB-36026: 
Cryo-EM structure of the Arabidopsis thaliana photosystem I in state 2(PSI-ST2)

EMDB-36032: 
LHCI of PSI from Arabidopsis thaliana in state 2 (PSI-ST2)

EMDB-36033: 
Cryo-EM structure of the Arabidopsis thaliana PSI in state 1 (PSI-ST1)

EMDB-36035: 
LHCI of PSI from Arabidopsis thaliana in state 1 (PSI-ST1)

EMDB-36036: 
Coordinates of Cryo-EM structure of the Arabidopsis thaliana PSI in state 1 (PSI-ST1)

EMDB-36037: 
Coordinates of Cryo-EM structure of the Arabidopsis thaliana PSI in state 2 (PSI-ST2)

EMDB-41075: 
SARS-CoV-2 spike in complex with Fab 71281-33

EMDB-41076: 
SARS-CoV-2 spike in complex with Fab 71281-33 (2)

EMDB-29281: 
Cryo-EM structure of STING oligomer bound to cGAMP and NVS-STG2

EMDB-29282: 
Cryo-EM structure of STING oligomer bound to cGAMP, NVS-STG2 and C53

EMDB-28793: 
Voltage-gated potassium channel Kv3.1 with novel positive modulator (9M)-9-{5-chloro-6-[(3,3-dimethyl-2,3-dihydro-1-benzofuran-4-yl)oxy]-4-methylpyridin-3-yl}-2-methyl-7,9-dihydro-8H-purin-8-one (compound 4)

EMDB-28795: 
Voltage-gated potassium channel Kv3.1 apo

EMDB-34475: 
Cryo-EM structure of the full transcription activation complex NtcA-NtcB-TAC

EMDB-34476: 
Cryo-EM structure of the transcription activation complex NtcA-TAC

EMDB-34477: 
Cryo-EM structure of the full transcription activation complex NtcA-NtcB-TAC focusing on NtcA and NtcB binding sites
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
