-Search query
-Search result
Showing 1 - 50 of 76 items for (author: chase & j)

EMDB-43762: 
Aca2 from Pectobacterium phage ZF40 bound to RNA

PDB-8w35: 
Aca2 from Pectobacterium phage ZF40 bound to RNA

EMDB-42681: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand

EMDB-42682: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the upper strand

EMDB-42683: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the upper strand

EMDB-42800: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the lower strand

EMDB-42833: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the lower strand

EMDB-42835: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the lower strand

EMDB-42846: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state from the upper strand

EMDB-42847: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the upper strand

EMDB-42849: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 1 from the lower strand

EMDB-42856: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 2 from the lower strand

EMDB-42858: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the lower strand

EMDB-42874: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand activated by the C1-domain of cardiac myosin binding protein C

PDB-8uww: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand

PDB-8uwx: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the upper strand

PDB-8uwy: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the upper strand

PDB-8uyd: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the lower strand

PDB-8uz5: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the lower strand

PDB-8uz6: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the lower strand

PDB-8uzx: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state from the upper strand

PDB-8uzy: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the upper strand

PDB-8v01: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 1 from the lower strand

PDB-8v0i: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 2 from the lower strand

PDB-8v0k: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the lower strand

PDB-8v0y: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand activated by the C1-domain of cardiac myosin binding protein C
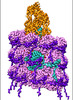
EMDB-15345: 
CryoEM structure of sweet potato feathery mottle virus VLP
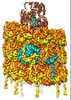
EMDB-15346: 
CryoEM structure of sweet potato mild mottle virus VLP

PDB-8acb: 
CryoEM structure of sweet potato feathery mottle virus VLP

PDB-8acc: 
CryoEM structure of sweet potato mild mottle virus VLP

EMDB-27178: 
Structure of Cas12a2 binary complex
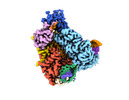
EMDB-27179: 
Structure of Cas12a2 ternary complex
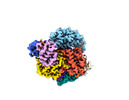
EMDB-27180: 
Structure of Cas12a2 ternary complex

PDB-8d49: 
Structure of Cas12a2 binary complex

PDB-8d4a: 
Cas12a2 quaternary complex

PDB-8d4b: 
Structure of Cas12a2 ternary complex
EMDB-27331: 
The structure of the native cardiac thin filament junction region

PDB-8dd0: 
The structure of the native cardiac thin filament junction region

EMDB-28089: 
a22L prion fibril
PDB-8efu: 
a22L prion fibril

EMDB-32331: 
Cryo-EM structure of the alpha2A adrenergic receptor GoA signaling complex bound to a G protein biased agonist

EMDB-32342: 
Cryo-EM structure of the alpha2A adrenergic receptor GoA signaling complex bound to a biased agonist

PDB-7w6p: 
Cryo-EM structure of the alpha2A adrenergic receptor GoA signaling complex bound to a G protein biased agonist

PDB-7w7e: 
Cryo-EM structure of the alpha2A adrenergic receptor GoA signaling complex bound to a biased agonist

EMDB-24252: 
Cryo-EM structure of MFSD2A

PDB-7n98: 
Cryo-EM structure of MFSD2A

EMDB-22964: 
Structure of cardiac native thin filament at pCa=5.8 having upper and lower troponins in Ca2+ free state

EMDB-22965: 
Structure of cardiac native thin filament at pCa=5.8 having upper and lower troponins in Ca2+ bound state

EMDB-22966: 
Structure of the native cardiac thin filament at pCa=5.8 having upper Tn in Ca2+ free state and lower Tn in Ca2+ bound state

EMDB-22978: 
Structure of upper Tn Ca2+ free (rotated) and lower Tn Ca2+ bound cardiac native thin filament at pCa=5.8
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
