-Search query
-Search result
Showing 1 - 50 of 723 items for (author: cao & sh)

EMDB-37546: 
Spike Trimer of BA.2.86 in complex with one hACE2
Method: single particle / : Yue C, Liu P

EMDB-37548: 
Spike Trimer of BA.2.86 in complex with two hACE2s
Method: single particle / : Yue C, Liu P

EMDB-37549: 
Spike Trimer of BA.2.86 with three RBDs down
Method: single particle / : Yue C, Liu P

EMDB-37550: 
Spike Trimer of BA.2.86 with single RBD up
Method: single particle / : Yue C, Liu P

EMDB-38049: 
SARS-CoV-2 JN.1 Spike
Method: single particle / : Yue C, Liu P

EMDB-38072: 
SARS-CoV-2 BA.2.75 Spike with K356T mutation (3 RBD down)
Method: single particle / : Yue C, Liu P

EMDB-38073: 
SARS-CoV-2 BA.2.75 Spike with K356T mutation (1 RBD up)
Method: single particle / : Yue C, Liu P

EMDB-38681: 
BA.2.86 Spike Trimer in complex with heparan sulfate
Method: single particle / : Yue C, Liu P

EMDB-38682: 
JN.1 Spike Trimer in complex with heparan sulfate
Method: single particle / : Yue C, Liu P

EMDB-38683: 
XBB.1.5 Spike Trimer in complex with heparan sulfate
Method: single particle / : Yue C, Liu P, Mao X

EMDB-38684: 
BA.2.86-T356K Spike Trimer in complex with heparan sulfate (Local refinement)
Method: single particle / : Yue C, Liu P

EMDB-37553: 
BA.2.86 RBD in complex with hACE2 (local refinement)
Method: single particle / : Yue C, Liu P

EMDB-38056: 
BA.2.86 Spike Trimer with ins483V mutation (3 RBD down)
Method: single particle / : Yue C, Liu P

EMDB-38057: 
BA.2.86 Spike Trimer with ins483V mutation (1 RBD up)
Method: single particle / : Yue C, Liu P

EMDB-38063: 
BA.2.86 Spike Trimer with T356K mutation (3 RBD down)
Method: single particle / : Yue C, Liu P

EMDB-38064: 
BA.2.86 Spike Trimer with T356K mutation (1 RBD up)
Method: single particle / : Yue C, Liu P

EMDB-38700: 
XBB.1.5-K356T S-trimer (1 RBD up)
Method: single particle / : Yue C, Liu P

EMDB-38701: 
XBB.1.5-K356T S-trimer (3 RBDs down)
Method: single particle / : Yue C, Liu P

EMDB-28966: 
CryoEM map of de novo designed oligomeric protein C4-71_6x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28967: 
CryoEM map of de novo designed oligomeric protein C4-71_8x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28968: 
CryoEM map of de novo designed oligomeric protein C6-71
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28969: 
CryoEM map of de novo designed oligomeric protein C6-71_6x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28970: 
CryoEM map of de novo designed oligomeric protein C6-71_8x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28971: 
CryoEM map of de novo designed oligomeric protein C8-71_6x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28972: 
CryoEM map of de novo designed oligomeric protein C8-71_8x
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28973: 
CryoEM map of de novo designed oligomeric protein C4-81
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-28974: 
CryoEM map of designed oligomeric protein C4-71
Method: single particle / : Redler RL, Edman NI, Baker D, Ekiert DC, Bhabha G

EMDB-38679: 
Cryo-EM structure of tomato NRC2 dimer
Method: single particle / : Sun Y, Ma SC, Chai JJ

EMDB-38680: 
Cryo-EM structure of tomato NRC2 tetramer
Method: single particle / : Sun Y, Ma SC, Chai JJ

EMDB-38685: 
Cryo-EM structure of tomato NRC2 filament
Method: helical / : Sun Y, Ma SC, Chai JJ

PDB-8xuq: 
Cryo-EM structure of tomato NRC2 tetramer
Method: single particle / : Sun Y, Ma SC, Chai JJ

EMDB-38394: 
The Cryo-EM structure of MPXV E5 apo conformation
Method: single particle / : Zhang W, Liu Y, Gao H, Gan J

EMDB-38395: 
The Cryo-EM structure of MPXV E5 in complex with DNA
Method: single particle / : Zhang W, Liu Y, Gao H, Gan J

EMDB-38396: 
The Cryo-EM structure of MPXV E5 C-terminal in complex with DNA
Method: single particle / : Zhang W, Liu Y, Gao H, Gan J

PDB-8xj6: 
The Cryo-EM structure of MPXV E5 apo conformation
Method: single particle / : Zhang W, Liu Y, Gao H, Gan J

PDB-8xj7: 
The Cryo-EM structure of MPXV E5 in complex with DNA
Method: single particle / : Zhang W, Liu Y, Gao H, Gan J

PDB-8xj8: 
The Cryo-EM structure of MPXV E5 C-terminal in complex with DNA
Method: single particle / : Zhang W, Liu Y, Gao H, Gan J
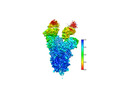
EMDB-37240: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Method: single particle / : Cao L, Wang X

EMDB-37241: 
The interface structure of Omicron RBD binding to 5817 Fab
Method: single particle / : Cao L, Wang X

PDB-8khc: 
SARS-CoV-2 Omicron spike in complex with 5817 Fab
Method: single particle / : Cao L, Wang X

PDB-8khd: 
The interface structure of Omicron RBD binding to 5817 Fab
Method: single particle / : Cao L, Wang X
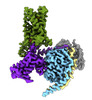
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL
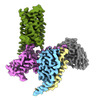
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)
Method: single particle / : Kim Y, Gumpper RH, Roth BL

PDB-8vy7: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

PDB-8vy9: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant
Method: single particle / : Kim Y, Gumpper RH, Roth BL

EMDB-36870: 
Structure of full Banna virus
Method: single particle / : Li Z, Cao S
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
