-Search query
-Search result
Showing 1 - 50 of 159 items for (author: blaza & j)

EMDB-18049: 
Chlorella sorokiniana Rubisco: D4 symmetry imposed
Method: single particle / : Barrett J, Blaza JN, Mackinder LCM

EMDB-18050: 
Chlorella sorokiniana Rubisco with CsLinker (alpha3-alpha4) bound: D4 symmetry expanded
Method: single particle / : Barrett J, Blaza JN, Mackinder LCM

PDB-8q04: 
Chlorella sorokiniana Rubisco: D4 symmetry imposed
Method: single particle / : Barrett J, Blaza JN, Mackinder LCM

PDB-8q05: 
Chlorella sorokiniana Rubisco with CsLinker (alpha3-alpha4) bound: D4 symmetry expanded
Method: single particle / : Barrett J, Blaza JN, Mackinder LCM
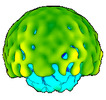
EMDB-50358: 
In vitro-induced genome-releasing intermediate of Rhodobacter microvirus Ebor computed with C5 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

EMDB-50356: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Byrom L, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

EMDB-50357: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA
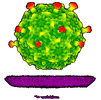
EMDB-50359: 
Rhodobacter microvirus Ebor attached to B10 host cell reconstructed by single particle analysis with applied C5 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA
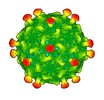
EMDB-50360: 
Rhodobacter microvirus Ebor attached to the outer membrane vesicle
Method: subtomogram averaging / : Bardy P, Blaza JN, Jenkins HT, Nicholas TR, Konig HC, Alim NTB, Hart SJ, Turkenburg JP, Fogg PCM, Beatty JT, Antson AA

EMDB-50361: 
Rhodobacter microvirus Ebor attached to the host cell reconstructed by subtomogram averaging
Method: subtomogram averaging / : Bardy P, Traore DAK, Blaza JN, Jenkins HT, Nicholas TR, Hart SJ, Turkenburg JP, Fogg PCM, Antson AA

PDB-9ffg: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Byrom L, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

PDB-9ffh: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
Method: single particle / : Bardy P, MacDonald CIW, Jenkins HT, Chechik M, Hart SJ, Turkenburg JP, Blaza JN, Fogg PCM, Antson AA

EMDB-19408: 
Cryo-EM structure of CDK2-cyclin A in complex with CDC25A
Method: single particle / : Rowland RJ, Noble MEM, Endicott JA

EMDB-19067: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).
Method: single particle / : Helena-Bueno K, Rybak MY, Gagnon MG, Hill CH, Melnikov SV

EMDB-19076: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).
Method: single particle / : Helena-Bueno K, Rybak MY, Gagnon MG, Hill CH, Melnikov SV

EMDB-19077: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).
Method: single particle / : Helena-Bueno K, Rybak MY, Gagnon MG, Hill CH, Melnikov SV

PDB-8rd8: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factors Balon and RaiA (structure 1).
Method: single particle / : Helena-Bueno K, Rybak MY, Gagnon MG, Hill CH, Melnikov SV

PDB-8rdv: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon, mRNA and P-site tRNA (structure 2).
Method: single particle / : Helena-Bueno K, Rybak MY, Gagnon MG, Hill CH, Melnikov SV

PDB-8rdw: 
Cryo-EM structure of P. urativorans 70S ribosome in complex with hibernation factor Balon and EF-Tu(GDP) (structure 3).
Method: single particle / : Helena-Bueno K, Rybak MY, Gagnon MG, Hill CH, Melnikov SV
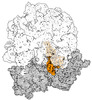
EMDB-43074: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) (Structure 4)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG
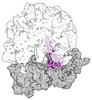
EMDB-43075: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Rv2629 (Balon) (Structure 5)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG
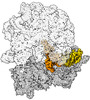
EMDB-43076: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) (Composite structure 6)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG

EMDB-43077: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) (Structure 6)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG

EMDB-43078: 
Hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) bound to Mycobacterium smegmatis 70S ribosome, from focused 3D classification and refinement (Structure 6)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG

PDB-8v9j: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) (Structure 4)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG

PDB-8v9k: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Rv2629 (Balon) (Structure 5)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG

PDB-8v9l: 
Cryo-EM structure of the Mycobacterium smegmatis 70S ribosome in complex with hibernation factor Msmeg1130 (Balon) and MsmegEF-Tu(GDP) (Composite structure 6)
Method: single particle / : Rybak MY, Helena-Bueno K, Hill CH, Melnikov SV, Gagnon MG

EMDB-16433: 
Cryo-EM structure of NADH bound SLA dehydrogenase RlGabD from Rhizobium leguminosarum bv. trifolii SRD1565
Method: single particle / : Sharma M, Meek RW, Armstrong Z, Blaza JN, Alhifthi A, Li J, Goddard-Borger ED, Williams SJ, Davies GJ

PDB-8c54: 
Cryo-EM structure of NADH bound SLA dehydrogenase RlGabD from Rhizobium leguminosarum bv. trifolii SRD1565
Method: single particle / : Sharma M, Meek RW, Armstrong Z, Blaza JN, Alhifthi A, Li J, Goddard-Borger ED, Williams SJ, Davies GJ

EMDB-16325: 
Cryo-EM structure of SKP1-SKP2-CKS1-CDK2-CyclinA-p27KIP1 Complex
Method: single particle / : Rowland RJ, Salamina M, Endicott JA, Noble ME
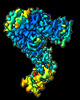
EMDB-16327: 
Cryo-EM structure of SKP1-SKP2-CKS1 from the SCFSKP2 E3 ligase complex
Method: single particle / : Rowland RJ, Salamina M, Endicott JA, Noble MEM
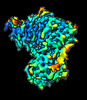
EMDB-16344: 
Cryo-EM structure of CDK2-CyclinA in complex with p27 from the SCFSKP2 E3 ligase Complex
Method: single particle / : Rowland RJ, Salamina M, Endicott JA, Noble ME

PDB-8bya: 
Cryo-EM structure of SKP1-SKP2-CKS1-CDK2-CyclinA-p27KIP1 Complex
Method: single particle / : Rowland RJ, Salamina M, Endicott JA, Noble ME

PDB-8byl: 
Cryo-EM structure of SKP1-SKP2-CKS1 from the SCFSKP2 E3 ligase complex
Method: single particle / : Rowland RJ, Salamina M, Endicott JA, Noble MEM

PDB-8bzo: 
Cryo-EM structure of CDK2-CyclinA in complex with p27 from the SCFSKP2 E3 ligase Complex
Method: single particle / : Rowland RJ, Salamina M, Endicott JA, Noble ME

EMDB-14126: 
Bovine complex I in the slack state at 3.5 A
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14128: 
Bovine complex I in the active state at 3.1 A (Composite map)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14129: 
Bovine complex I in the active state at 3.1 A (Hydrophlic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14130: 
Bovine complex I in the active state at 3.1 A (Proximal hydrophobic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14131: 
Bovine complex I in the active state at 3.1 A (Distal hydrophobic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14251: 
Bovine complex I in the presence of IM1761092, active class i (Composite map)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14252: 
Bovine complex I in the presence of IM1761092, active class i (Consensus)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14253: 
Bovine complex I in the presence of IM1761092, active class i (Hydophilic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14254: 
Bovine complex I in the presence of IM1761092, active class i (Proximal hydrophobic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14255: 
Bovine complex I in the presence of IM1761092, active class i (Distal hydrophobic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14256: 
Bovine complex I in the presence of IM1761092, active class ii (Composite map)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14257: 
Bovine complex I in the presence of IM1761092, active class ii (Consensus map)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14258: 
Bovine complex I in the presence of IM1761092, active class ii (Hydrophilic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14259: 
Bovine complex I in the presence of IM1761092, active class ii (Proximal hydrophobic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J

EMDB-14260: 
Bovine complex I in the presence of IM1761092, active class ii (Distal hydrophobic domain)
Method: single particle / : Bridges HR, Blaza JN, Yin Z, Chung I, Hirst J
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
