[English] 日本語
 Yorodumi
Yorodumi- EMDB-16327: Cryo-EM structure of SKP1-SKP2-CKS1 from the SCFSKP2 E3 ligase complex -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of SKP1-SKP2-CKS1 from the SCFSKP2 E3 ligase complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | cell cycle / cyclin-dependent kinase / signalling / ubiquitinationCell cycle | |||||||||
| Function / homology |  Function and homology information Function and homology informationcyclin-dependent protein kinase regulator activity / regulation of lens fiber cell differentiation / negative regulation of cardiac muscle tissue regeneration / positive regulation of protein polyubiquitination / autophagic cell death / FOXO-mediated transcription of cell cycle genes / negative regulation of epithelial cell proliferation involved in prostate gland development / cellular response to cell-matrix adhesion / F-box domain binding / Aberrant regulation of mitotic exit in cancer due to RB1 defects ...cyclin-dependent protein kinase regulator activity / regulation of lens fiber cell differentiation / negative regulation of cardiac muscle tissue regeneration / positive regulation of protein polyubiquitination / autophagic cell death / FOXO-mediated transcription of cell cycle genes / negative regulation of epithelial cell proliferation involved in prostate gland development / cellular response to cell-matrix adhesion / F-box domain binding / Aberrant regulation of mitotic exit in cancer due to RB1 defects / regulation of cell cycle G1/S phase transition / PcG protein complex / epithelial cell proliferation involved in prostate gland development / regulation of exit from mitosis / negative regulation of epithelial cell apoptotic process / cyclin-dependent protein serine/threonine kinase inhibitor activity / maintenance of protein location in nucleus / Cul7-RING ubiquitin ligase complex / RHO GTPases activate CIT / regulation of cyclin-dependent protein serine/threonine kinase activity / Loss of Function of FBXW7 in Cancer and NOTCH1 Signaling / nuclear export / cyclin-dependent protein serine/threonine kinase activator activity / negative regulation of mitotic cell cycle / epithelial cell apoptotic process / AKT phosphorylates targets in the cytosol / ubiquitin ligase activator activity / molecular function inhibitor activity / cellular response to lithium ion / positive regulation of intracellular estrogen receptor signaling pathway / SCF ubiquitin ligase complex / p53-Dependent G1 DNA Damage Response / PTK6 Regulates Cell Cycle / SCF-dependent proteasomal ubiquitin-dependent protein catabolic process / Cul4A-RING E3 ubiquitin ligase complex / Constitutive Signaling by AKT1 E17K in Cancer / Prolactin receptor signaling / cellular response to antibiotic / Defective binding of RB1 mutants to E2F1,(E2F2, E2F3) / inner ear development / ciliary transition zone / ubiquitin ligase complex scaffold activity / protein kinase inhibitor activity / regulation of G1/S transition of mitotic cell cycle / negative regulation of vascular associated smooth muscle cell proliferation / Estrogen-dependent nuclear events downstream of ESR-membrane signaling / cullin family protein binding / TP53 Regulates Transcription of Genes Involved in G1 Cell Cycle Arrest / protein monoubiquitination / protein K63-linked ubiquitination / Cyclin E associated events during G1/S transition / Cyclin A:Cdk2-associated events at S phase entry / cyclin-dependent protein kinase holoenzyme complex / positive regulation of double-strand break repair via homologous recombination / positive regulation of microtubule polymerization / ubiquitin-like ligase-substrate adaptor activity / Notch signaling pathway / Nuclear events stimulated by ALK signaling in cancer / protein K48-linked ubiquitination / FLT3 Signaling / regulation of mitotic cell cycle / positive regulation of smooth muscle cell proliferation / regulation of cell migration / cyclin binding / molecular function activator activity / placenta development / positive regulation of DNA replication / Regulation of BACH1 activity / MAP3K8 (TPL2)-dependent MAPK1/3 activation / ubiquitin binding / DNA damage response, signal transduction by p53 class mediator / potassium ion transport / SCF-beta-TrCP mediated degradation of Emi1 / NIK-->noncanonical NF-kB signaling / Vpu mediated degradation of CD4 / Dectin-1 mediated noncanonical NF-kB signaling / Degradation of CRY and PER proteins / sensory perception of sound / G1/S transition of mitotic cell cycle / Activation of NF-kappaB in B cells / Iron uptake and transport / Degradation of GLI1 by the proteasome / GSK3B and BTRC:CUL1-mediated-degradation of NFE2L2 / Negative regulation of NOTCH4 signaling / negative regulation of cell growth / APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 / FBXL7 down-regulates AURKA during mitotic entry and in early mitosis / Degradation of GLI2 by the proteasome / GLI3 is processed to GLI3R by the proteasome / Ubiquitin-Mediated Degradation of Phosphorylated Cdc25A / beta-catenin binding / DNA Damage/Telomere Stress Induced Senescence / NOTCH1 Intracellular Domain Regulates Transcription / Degradation of beta-catenin by the destruction complex / G2/M transition of mitotic cell cycle / CLEC7A (Dectin-1) signaling / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants / SCF(Skp2)-mediated degradation of p27/p21 / FCERI mediated NF-kB activation Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Rowland RJ / Salamina M / Endicott JA / Noble MEM | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Sci Rep / Year: 2023 Journal: Sci Rep / Year: 2023Title: Cryo-EM structure of SKP1-SKP2-CKS1 in complex with CDK2-cyclin A-p27KIP1. Authors: Rhianna J Rowland / Richard Heath / Daniel Maskell / Rebecca F Thompson / Neil A Ranson / James N Blaza / Jane A Endicott / Martin E M Noble / Marco Salamina /  Abstract: p27KIP1 (cyclin-dependent kinase inhibitor 1B, p27) is a member of the CIP/KIP family of CDK (cyclin dependent kinase) regulators that inhibit cell cycle CDKs. p27 phosphorylation by CDK1/2, signals ...p27KIP1 (cyclin-dependent kinase inhibitor 1B, p27) is a member of the CIP/KIP family of CDK (cyclin dependent kinase) regulators that inhibit cell cycle CDKs. p27 phosphorylation by CDK1/2, signals its recruitment to the SCF (S-phase kinase associated protein 1 (SKP1)-cullin-SKP2) E3 ubiquitin ligase complex for proteasomal degradation. The nature of p27 binding to SKP2 and CKS1 was revealed by the SKP1-SKP2-CKS1-p27 phosphopeptide crystal structure. Subsequently, a model for the hexameric CDK2-cyclin A-CKS1-p27-SKP1-SKP2 complex was proposed by overlaying an independently determined CDK2-cyclin A-p27 structure. Here we describe the experimentally determined structure of the isolated CDK2-cyclin A-CKS1-p27-SKP1-SKP2 complex at 3.4 Å global resolution using cryogenic electron microscopy. This structure supports previous analysis in which p27 was found to be structurally dynamic, transitioning from disordered to nascent secondary structure on target binding. We employed 3D variability analysis to further explore the conformational space of the hexameric complex and uncovered a previously unidentified hinge motion centred on CKS1. This flexibility gives rise to open and closed conformations of the hexameric complex that we propose may contribute to p27 regulation by facilitating recognition with SCF. This 3D variability analysis further informed particle subtraction and local refinement approaches to enhance the local resolution of the complex. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16327.map.gz emd_16327.map.gz | 97.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16327-v30.xml emd-16327-v30.xml emd-16327.xml emd-16327.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
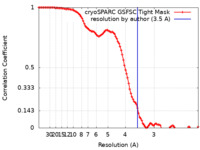
| FSC (resolution estimation) |  emd_16327_fsc.xml emd_16327_fsc.xml | 9.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_16327.png emd_16327.png | 116.9 KB | ||
| Masks |  emd_16327_msk_1.map emd_16327_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16327.cif.gz emd-16327.cif.gz | 6.6 KB | ||
| Others |  emd_16327_half_map_1.map.gz emd_16327_half_map_1.map.gz emd_16327_half_map_2.map.gz emd_16327_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16327 http://ftp.pdbj.org/pub/emdb/structures/EMD-16327 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16327 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16327 | HTTPS FTP |
-Related structure data
| Related structure data |  8bylMC  8byaC  8bzoC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16327.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16327.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||
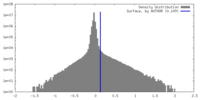
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16327_msk_1.map emd_16327_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
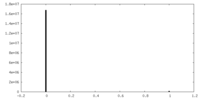
| Density Histograms |
-Half map: #1
| File | emd_16327_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
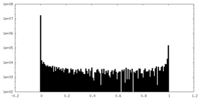
| Density Histograms |
-Half map: #2
| File | emd_16327_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
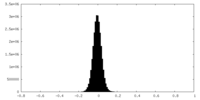
| Density Histograms |
- Sample components
Sample components
-Entire : Locally refined complex of SKP1-SKP2-CKS1-p27 from the hexametric...
| Entire | Name: Locally refined complex of SKP1-SKP2-CKS1-p27 from the hexametric SCFSKP2 E3 Ligase complex |
|---|---|
| Components |
|
-Supramolecule #1: Locally refined complex of SKP1-SKP2-CKS1-p27 from the hexametric...
| Supramolecule | Name: Locally refined complex of SKP1-SKP2-CKS1-p27 from the hexametric SCFSKP2 E3 Ligase complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 66 KDa |
-Macromolecule #1: S-phase kinase-associated protein 1
| Macromolecule | Name: S-phase kinase-associated protein 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.679965 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPSIKLQSSD GEIFEVDVEI AKQSVTIKTM LEDLGMDDEG DDDPVPLPNV NAAILKKVIQ WCTHHKDDPP PPEDDENKEK RTDDIPVWD QEFLKVDQGT LFELILAANY LDIKGLLDVT CKTVANMIKG KTPEEIRKTF NIKNDFTEEE EAQVRKENQW C EEK UniProtKB: S-phase kinase-associated protein 1 |
-Macromolecule #2: S-phase kinase-associated protein 2
| Macromolecule | Name: S-phase kinase-associated protein 2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.817785 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHRKHLQEIP DLSSNVATSF TWGWDSSKTS ELLSGMGVSA LEKEEPDSEN IPQELLSNLG HPESPPRKRL KSKGSDKDFV IVRRPKLNR ENFPGVSWDS LPDELLLGIF SCLCLPELLK VSGVCKRWYR LASDESLWQT LDLTGKNLHP DVTGRLLSQG V IAFRCPRS ...String: MHRKHLQEIP DLSSNVATSF TWGWDSSKTS ELLSGMGVSA LEKEEPDSEN IPQELLSNLG HPESPPRKRL KSKGSDKDFV IVRRPKLNR ENFPGVSWDS LPDELLLGIF SCLCLPELLK VSGVCKRWYR LASDESLWQT LDLTGKNLHP DVTGRLLSQG V IAFRCPRS FMDQPLAEHF SPFRVQHMDL SNSVIEVSTL HGILSQCSKL QNLSLEGLRL SDPIVNTLAK NSNLVRLNLS GC SGFSEFA LQTLLSSCSR LDELNLSWCF DFTEKHVQVA VAHVSETITQ LNLSGYRKNL QKSDLSTLVR RCPNLVHLDL SDS VMLKND CFQEFFQLNY LQHLSLSRCY DIIPETLLEL GEIPTLKTLQ VFGIVPDGTL QLLKEALPHL QINCSHFTTI ARPT IGNKK NQEIWGIKCR LTLQKPSCL UniProtKB: S-phase kinase-associated protein 2 |
-Macromolecule #3: Cyclin-dependent kinases regulatory subunit 1
| Macromolecule | Name: Cyclin-dependent kinases regulatory subunit 1 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 9.679211 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSHKQIYYSD KYDDEEFEYR HVMLPKDIAK LVPKTHLMSE SEWRNLGVQQ SQGWVHYMIH EPEPHILLFR RPLPKKPKK UniProtKB: Cyclin-dependent kinases regulatory subunit 1 |
-Macromolecule #4: Cyclin-dependent kinase inhibitor 1B
| Macromolecule | Name: Cyclin-dependent kinase inhibitor 1B / type: protein_or_peptide / ID: 4 / Details: DGSPNAGSVEQ(TPO)PKK / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 22.188303 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSNVRVSNGS PSLERMDARQ AEHPKPSACR NLFGPVDHEE LTRDLEKHCR DMEEASQRKW NFDFQNHKPL EGKYEWQEVE KGSLPEFYY RPPRPPKGAC KVPAQESQDV SGSRPAAPLI GAPANSEDTH LVDPKTDPSD SQTGLAEQCA GIRKRPATDD S STQNKRAN ...String: MSNVRVSNGS PSLERMDARQ AEHPKPSACR NLFGPVDHEE LTRDLEKHCR DMEEASQRKW NFDFQNHKPL EGKYEWQEVE KGSLPEFYY RPPRPPKGAC KVPAQESQDV SGSRPAAPLI GAPANSEDTH LVDPKTDPSD SQTGLAEQCA GIRKRPATDD S STQNKRAN RTEENVSDGS PNAGSVEQ(TPO)P KKPGLRRRQT UniProtKB: Cyclin-dependent kinase inhibitor 1B |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 278.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average exposure time: 9.0 sec. / Average electron dose: 65.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Initial fitting was performed in chimera followed by real space refinement in Phenix |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Overall B value: 122 |
| Output model |  PDB-8byl: |
 Movie
Movie Controller
Controller



























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)