-Search query
-Search result
Showing all 40 items for (author: battisti & a)

EMDB-5448: 
Cryo-tomography and subtomogram averaging of the Newcastle disease virus matrix protein array
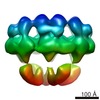
EMDB-2056: 
Electron cryo-microscopy of the Hantaan virus glycoprotein spike

EMDB-5106: 
Fab from MAb B interacting with feline panleukopenia virus

EMDB-5107: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 6

EMDB-5108: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 8

EMDB-5109: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 15

EMDB-5110: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 16

EMDB-5111: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb E

EMDB-5112: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb F

EMDB-1597: 
5-fold averaged density map of Paramecium bursaria Chlorella virus-1 (PBCV-1).

PDB-3iy0: 
Variable domains of the x-ray structure of Fab 14 fitted into the cryoEM reconstruction of the virus-Fab 14 complex

PDB-3iy1: 
Variable domains of the WAM of Fab B fitted into the cryoEM reconstruction of the virus-Fab B complex

PDB-3iy2: 
Variable domains of the computer generated model (WAM) of Fab 6 fitted into the cryoEM reconstruction of the virus-Fab 6 complex

PDB-3iy3: 
Variable domains of the computer generated model (WAM) of Fab 8 fitted into the cryoEM reconstruction of the virus-Fab 8 complex

PDB-3iy4: 
Variable domains of the computer generated model (WAM) of Fab 15 fitted into the cryoEM reconstruction of the virus-Fab 15 complex

PDB-3iy5: 
Variable domains of the mouse Fab (1AIF) fitted into the cryoEM reconstruction of the virus-Fab 16 complex

PDB-3iy6: 
Variable domains of the computer generated model (WAM) of Fab E fitted into the cryoEM reconstruction of the virus-Fab E complex

PDB-3iy7: 
Variable domains of the computer generated model (WAM) of Fab F fitted into the cryoEM reconstruction of the virus-Fab F complex

EMDB-5105: 
Canine parvovirus in complex with FAb from neutralizing antibody MAb 14

EMDB-1580: 
CryoEM and 3D image reconstruction of Chilo Iridescent virus at 13 angstrom resolution

EMDB-1418: 
Binding of a neutralizing antibody to dengue virus alters the arrangement of surface glycoproteins

PDB-2r6p: 
Fit of E protein and Fab 1A1D-2 into 24 angstrom resolution cryoEM map of Fab complexed with dengue 2 virus.

EMDB-1280: 
Cryo-electron microscopy study of bacteriophage T4 displaying anthrax toxin proteins.

EMDB-1266: 
Structural changes of bacteriophage phi29 upon DNA packaging and release.

EMDB-1393: 
Cryo-EM study of the Pseudomonas bacteriophage phiKZ.

EMDB-1392: 
A three-dimensional cryo-electron microscopy structure of the bacteriophage phiKZ head.

EMDB-1415: 
Cryo-EM study of the Pseudomonas bacteriophage phiKZ.

EMDB-1333: 
The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities.

EMDB-1334: 
The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities.

EMDB-1335: 
The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities.

EMDB-1336: 
The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities.

EMDB-1337: 
The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities.

PDB-2nsu: 
Crystal structure of the ectodomain of human transferrin receptor fitted into a cryo-EM reconstruction of canine parvovirus and feline transferrin receptor complex

EMDB-1287: 
Asymmetric binding of transferrin receptor to parvovirus capsids.

EMDB-1288: 
Asymmetric binding of transferrin receptor to parvovirus capsids.

EMDB-1260: 
Structural changes of bacteriophage phi29 upon DNA packaging and release.

EMDB-1265: 
Structural changes of bacteriophage phi29 upon DNA packaging and release.

EMDB-1166: 
Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN.
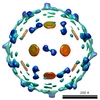
EMDB-1167: 
Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN.

PDB-2b6b: 
Cryo EM structure of Dengue complexed with CRD of DC-SIGN
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
