-Search query
-Search result
Showing 1 - 50 of 824 items for (author: ando & n)

EMDB-44635: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM

PDB-9bjk: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM

EMDB-43506: 
Cryo-EM structure of LKB1-STRADalpha-MO25alpha heterocomplex

PDB-8vsu: 
Cryo-EM structure of LKB1-STRADalpha-MO25alpha heterocomplex

EMDB-41433: 
Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter

EMDB-41437: 
Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter

EMDB-41439: 
Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter

EMDB-41448: 
Escherichia coli RNA polymerase unwinding intermediate (I1c) at the lambda PR promoter

EMDB-41456: 
Escherichia coli RNA polymerase closed complex intermediate at the lambda PR promoter

PDB-8to1: 
Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter

PDB-8to6: 
Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter

PDB-8to8: 
Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter

PDB-8toe: 
Escherichia coli RNA polymerase unwinding intermediate (I1c) at the lambda PR promoter

PDB-8tom: 
Escherichia coli RNA polymerase closed complex intermediate at the lambda PR promoter

EMDB-43527: 
Apoferritin at 100 keV on Alpine detector with a side-entry cryoholder

EMDB-43528: 
Aldolase at 100 keV on the Alpine detector with a side-entry cryoholder

EMDB-17295: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

PDB-8oyt: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

EMDB-16466: 
The structural architecture of alpha-synuclein oligomer

EMDB-16528: 
3D reconstruction of alpha-synuclein oligomer-PSMa3 complex

EMDB-19426: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api137

EMDB-19427: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation I

EMDB-19428: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation II

EMDB-19429: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation III

PDB-8rpy: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api137

PDB-8rpz: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation I

PDB-8rq0: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation II

PDB-8rq2: 
Escherichia coli 50S subunit in complex with the antimicrobial peptide Api88 - conformation III

EMDB-17296: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

PDB-8oyu: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation
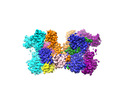
EMDB-43705: 
HIV-1 wild-type intasome core

EMDB-43756: 
HIV-1 P5-IN intasome core
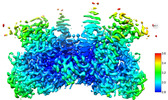
EMDB-43761: 
HIV-1 intasome core assembled with wild-type integrase, 1F

PDB-8w09: 
HIV-1 wild-type intasome core

PDB-8w2r: 
HIV-1 P5-IN intasome core

PDB-8w34: 
HIV-1 intasome core assembled with wild-type integrase, 1F

EMDB-41672: 
ELIC5 with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG

EMDB-41673: 
ELIC with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG

PDB-8twv: 
ELIC5 with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG

PDB-8twz: 
ELIC with Propylamine in spNW15 nanodiscs with 2:1:1 POPC:POPE:POPG

EMDB-18520: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

EMDB-18521: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

EMDB-18522: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

EMDB-18523: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

PDB-8qo2: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes
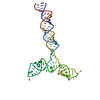
PDB-8qo3: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

PDB-8qo4: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

PDB-8qo5: 
Conserved Structures and Dynamics in 5-Proximal Regions of Betacoronavirus RNA Genomes

EMDB-43049: 
Cryo-electron tomogram of mitochondria from cryo-FIB milled l-Opa1* mouse embryonic fibroblasts

EMDB-43050: 
cryo-electron tomogram of mitochondria in cryo-FIB milled l-Opa1* mouse embryonic fibroblasts
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
